Magnesium »
PDB 1ao0-1bh2 »
1ba0 »
Magnesium in PDB 1ba0: Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3
Enzymatic activity of Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3
All present enzymatic activity of Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3:
3.6.1.3;
3.6.1.3;
Protein crystallography data
The structure of Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3, PDB code: 1ba0
was solved by
S.M.Wilbanks,
D.B.Mckay,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 8.00 / 1.90 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 143.300, 63.800, 46.500, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 21.6 / 27.5 |
Other elements in 1ba0:
The structure of Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3 also contains other interesting chemical elements:
| Chlorine | (Cl) | 2 atoms |
| Sodium | (Na) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3
(pdb code 1ba0). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3, PDB code: 1ba0:
In total only one binding site of Magnesium was determined in the Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3, PDB code: 1ba0:
Magnesium binding site 1 out of 1 in 1ba0
Go back to
Magnesium binding site 1 out
of 1 in the Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3
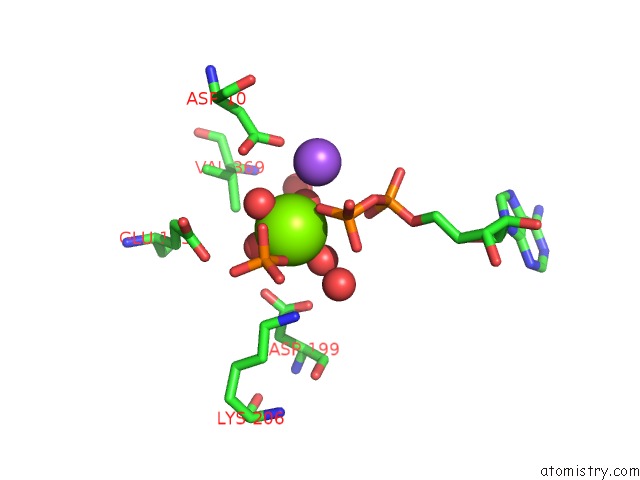
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Heat-Shock Cognate 70KD Protein 44KD Atpase N-Terminal 1NGE 3 within 5.0Å range:
|
Reference:
S.M.Wilbanks,
D.B.Mckay.
Structural Replacement of Active Site Monovalent Cations By the Epsilon-Amino Group of Lysine in the Atpase Fragment of Bovine HSC70. Biochemistry V. 37 7456 1998.
ISSN: ISSN 0006-2960
PubMed: 9585559
DOI: 10.1021/BI973046M
Page generated: Tue Aug 13 02:12:32 2024
ISSN: ISSN 0006-2960
PubMed: 9585559
DOI: 10.1021/BI973046M
Last articles
Cl in 5WHUCl in 5WKV
Cl in 5WKR
Cl in 5WKU
Cl in 5WKK
Cl in 5WKI
Cl in 5WJO
Cl in 5WKE
Cl in 5WKG
Cl in 5WHV