Magnesium »
PDB 1e9b-1enn »
1elx »
Magnesium in PDB 1elx: E. Coli Alkaline Phosphatase Mutant (S102A)
Enzymatic activity of E. Coli Alkaline Phosphatase Mutant (S102A)
All present enzymatic activity of E. Coli Alkaline Phosphatase Mutant (S102A):
3.1.3.1;
3.1.3.1;
Protein crystallography data
The structure of E. Coli Alkaline Phosphatase Mutant (S102A), PDB code: 1elx
was solved by
B.Stec,
M.Hehir,
C.Brennan,
M.Nolte,
E.R.Kantrowitz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 9.00 / 2.60 |
| Space group | P 63 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 163.620, 163.620, 138.880, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 15.8 / 20.7 |
Other elements in 1elx:
The structure of E. Coli Alkaline Phosphatase Mutant (S102A) also contains other interesting chemical elements:
| Zinc | (Zn) | 4 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the E. Coli Alkaline Phosphatase Mutant (S102A)
(pdb code 1elx). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the E. Coli Alkaline Phosphatase Mutant (S102A), PDB code: 1elx:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the E. Coli Alkaline Phosphatase Mutant (S102A), PDB code: 1elx:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 1elx
Go back to
Magnesium binding site 1 out
of 2 in the E. Coli Alkaline Phosphatase Mutant (S102A)
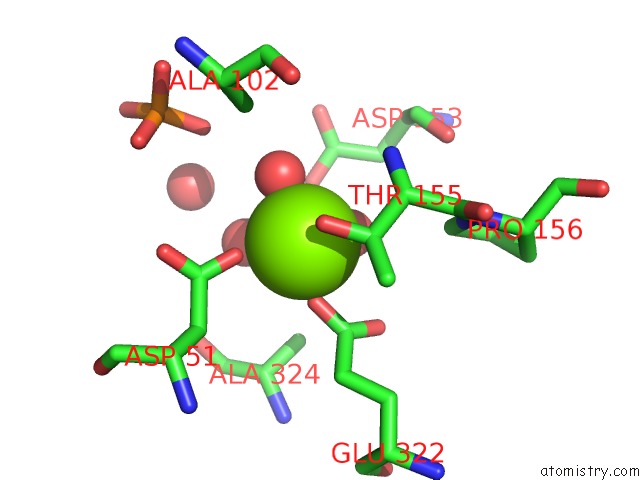
Mono view
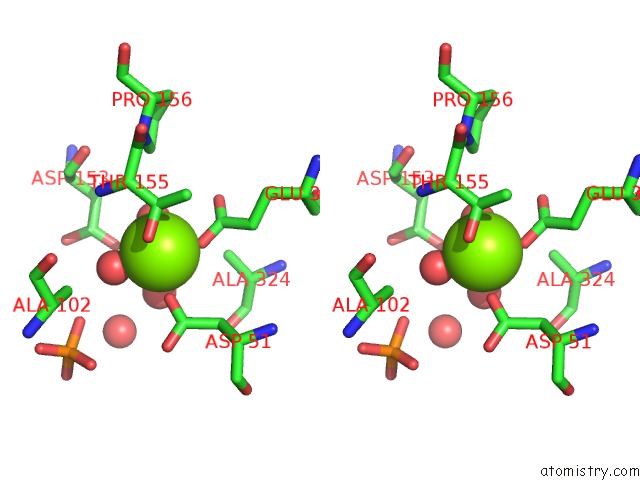
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of E. Coli Alkaline Phosphatase Mutant (S102A) within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 1elx
Go back to
Magnesium binding site 2 out
of 2 in the E. Coli Alkaline Phosphatase Mutant (S102A)
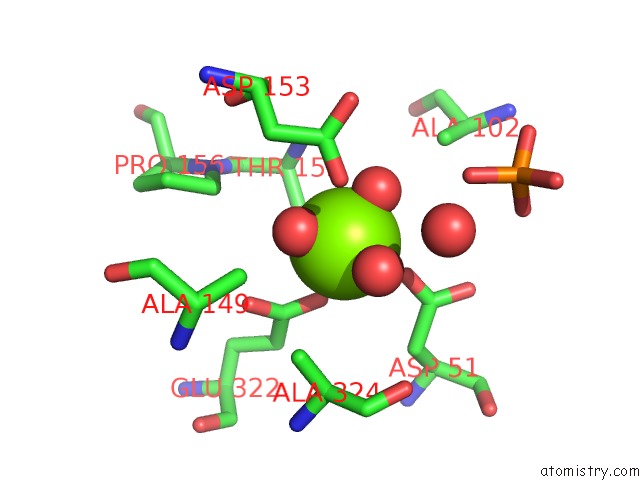
Mono view
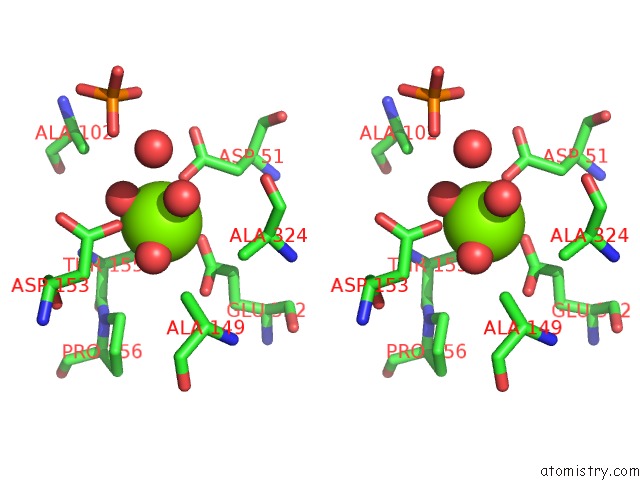
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of E. Coli Alkaline Phosphatase Mutant (S102A) within 5.0Å range:
|
Reference:
B.Stec,
M.J.Hehir,
C.Brennan,
M.Nolte,
E.R.Kantrowitz.
Kinetic and X-Ray Structural Studies of Three Mutant E. Coli Alkaline Phosphatases: Insights Into the Catalytic Mechanism Without the Nucleophile SER102. J.Mol.Biol. V. 277 647 1998.
ISSN: ISSN 0022-2836
PubMed: 9533886
DOI: 10.1006/JMBI.1998.1635
Page generated: Sat Aug 9 20:45:43 2025
ISSN: ISSN 0022-2836
PubMed: 9533886
DOI: 10.1006/JMBI.1998.1635
Last articles
Mg in 4DPGMg in 4DQP
Mg in 4DQQ
Mg in 4DPM
Mg in 4DPV
Mg in 4DQI
Mg in 4DOB
Mg in 4DOC
Mg in 4DMZ
Mg in 4DOA