Magnesium »
PDB 1so6-1t5s »
1sr6 »
Magnesium in PDB 1sr6: Structure of Nucleotide-Free Scallop Myosin S1
Protein crystallography data
The structure of Structure of Nucleotide-Free Scallop Myosin S1, PDB code: 1sr6
was solved by
D.Risal,
S.Gourinath,
D.M.Himmel,
A.G.Szent-Gyorgyi,
C.Cohen,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 38.79 / 2.75 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 83.591, 51.109, 162.329, 90.00, 98.02, 90.00 |
| R / Rfree (%) | 24.2 / 28.6 |
Other elements in 1sr6:
The structure of Structure of Nucleotide-Free Scallop Myosin S1 also contains other interesting chemical elements:
| Calcium | (Ca) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Structure of Nucleotide-Free Scallop Myosin S1
(pdb code 1sr6). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Structure of Nucleotide-Free Scallop Myosin S1, PDB code: 1sr6:
In total only one binding site of Magnesium was determined in the Structure of Nucleotide-Free Scallop Myosin S1, PDB code: 1sr6:
Magnesium binding site 1 out of 1 in 1sr6
Go back to
Magnesium binding site 1 out
of 1 in the Structure of Nucleotide-Free Scallop Myosin S1
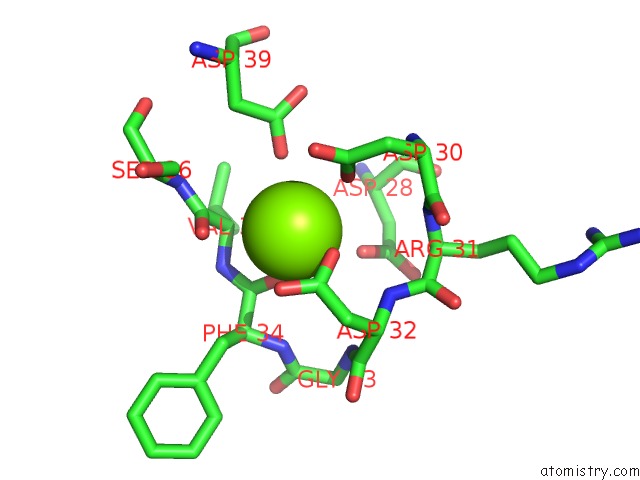
Mono view
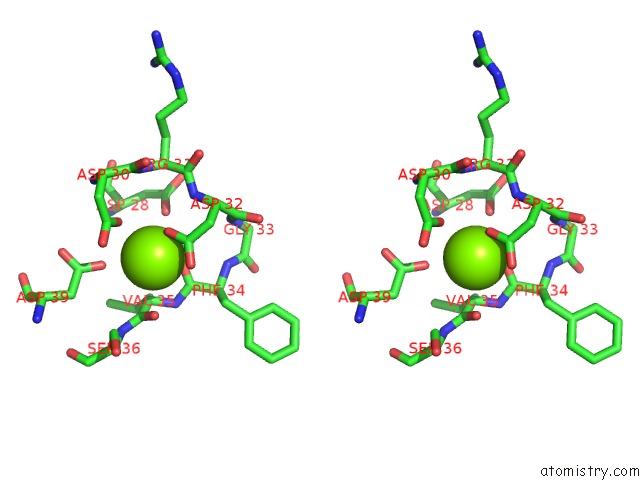
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Structure of Nucleotide-Free Scallop Myosin S1 within 5.0Å range:
|
Reference:
D.Risal,
S.Gourinath,
D.M.Himmel,
A.G.Szent-Gyorgyi,
C.Cohen.
Myosin Subfragment 1 Structures Reveal A Partially Bound Nucleotide and A Complex Salt Bridge That Helps Couple Nucleotide and Actin Binding. Proc.Natl.Acad.Sci.Usa V. 101 8930 2004.
ISSN: ISSN 0027-8424
PubMed: 15184651
DOI: 10.1073/PNAS.0403002101
Page generated: Sun Aug 10 04:42:58 2025
ISSN: ISSN 0027-8424
PubMed: 15184651
DOI: 10.1073/PNAS.0403002101
Last articles
Mg in 3G5AMg in 3G5S
Mg in 3G58
Mg in 3G4T
Mg in 3G4L
Mg in 3G4K
Mg in 3G4I
Mg in 3G4G
Mg in 3G4F
Mg in 3G45