Magnesium »
PDB 2f6y-2fl2 »
2f9g »
Magnesium in PDB 2f9g: Crystal Structure of FUS3 Phosphorylated on TYR182
Enzymatic activity of Crystal Structure of FUS3 Phosphorylated on TYR182
All present enzymatic activity of Crystal Structure of FUS3 Phosphorylated on TYR182:
2.7.1.37;
2.7.1.37;
Protein crystallography data
The structure of Crystal Structure of FUS3 Phosphorylated on TYR182, PDB code: 2f9g
was solved by
R.P.Bhattacharyya,
A.Remenyi,
M.C.Good,
C.J.Bashor,
A.M.Falick,
W.A.Lim,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 2.10 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 56.811, 62.533, 86.008, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 21.1 / 26 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of FUS3 Phosphorylated on TYR182
(pdb code 2f9g). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of FUS3 Phosphorylated on TYR182, PDB code: 2f9g:
In total only one binding site of Magnesium was determined in the Crystal Structure of FUS3 Phosphorylated on TYR182, PDB code: 2f9g:
Magnesium binding site 1 out of 1 in 2f9g
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of FUS3 Phosphorylated on TYR182
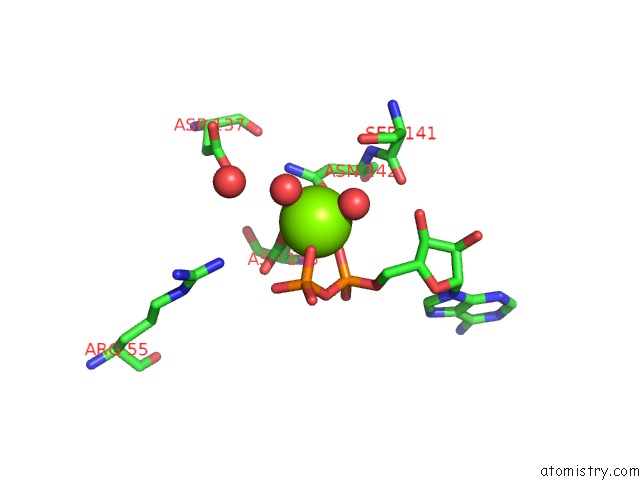
Mono view
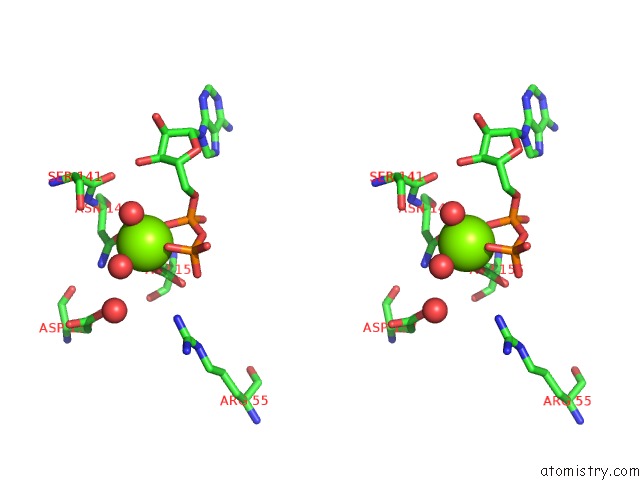
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of FUS3 Phosphorylated on TYR182 within 5.0Å range:
|
Reference:
R.P.Bhattacharyya,
A.Remenyi,
M.C.Good,
C.J.Bashor,
A.M.Falick,
W.A.Lim.
The STE5 Scaffold Allosterically Modulates Signaling Output of the Yeast Mating Pathway. Science V. 311 822 2006.
ISSN: ISSN 0036-8075
PubMed: 16424299
DOI: 10.1126/SCIENCE.1120941
Page generated: Tue Aug 13 23:05:15 2024
ISSN: ISSN 0036-8075
PubMed: 16424299
DOI: 10.1126/SCIENCE.1120941
Last articles
Ca in 5OSTCa in 5OSK
Ca in 5OLR
Ca in 5OLW
Ca in 5OP3
Ca in 5ONP
Ca in 5ONQ
Ca in 5OLS
Ca in 5OLP
Ca in 5OLQ