Magnesium »
PDB 2rex-2uxd »
2ukd »
Magnesium in PDB 2ukd: Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp
Enzymatic activity of Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp
All present enzymatic activity of Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp:
2.7.4.14;
2.7.4.14;
Protein crystallography data
The structure of Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp, PDB code: 2ukd
was solved by
I.Schlichting,
J.Reinstein,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 37.00 / 2.20 |
| Space group | P 41 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 78.700, 78.700, 100.100, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.3 / 23.1 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp
(pdb code 2ukd). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp, PDB code: 2ukd:
In total only one binding site of Magnesium was determined in the Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp, PDB code: 2ukd:
Magnesium binding site 1 out of 1 in 2ukd
Go back to
Magnesium binding site 1 out
of 1 in the Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp
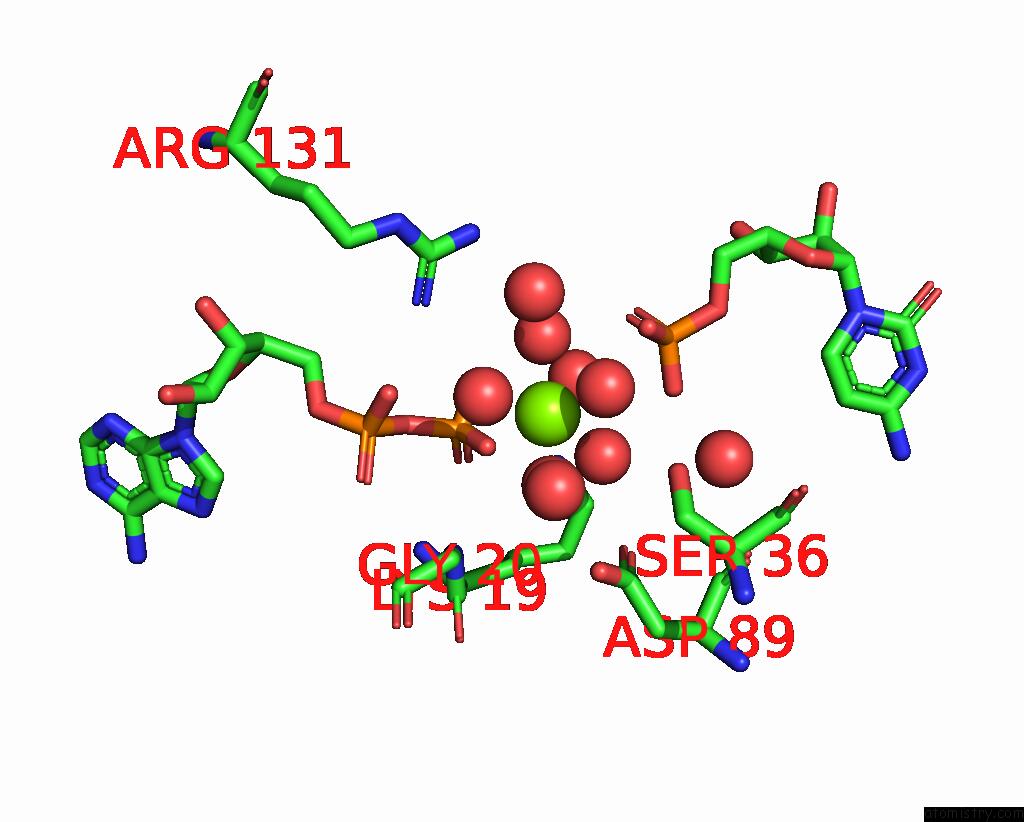
Mono view
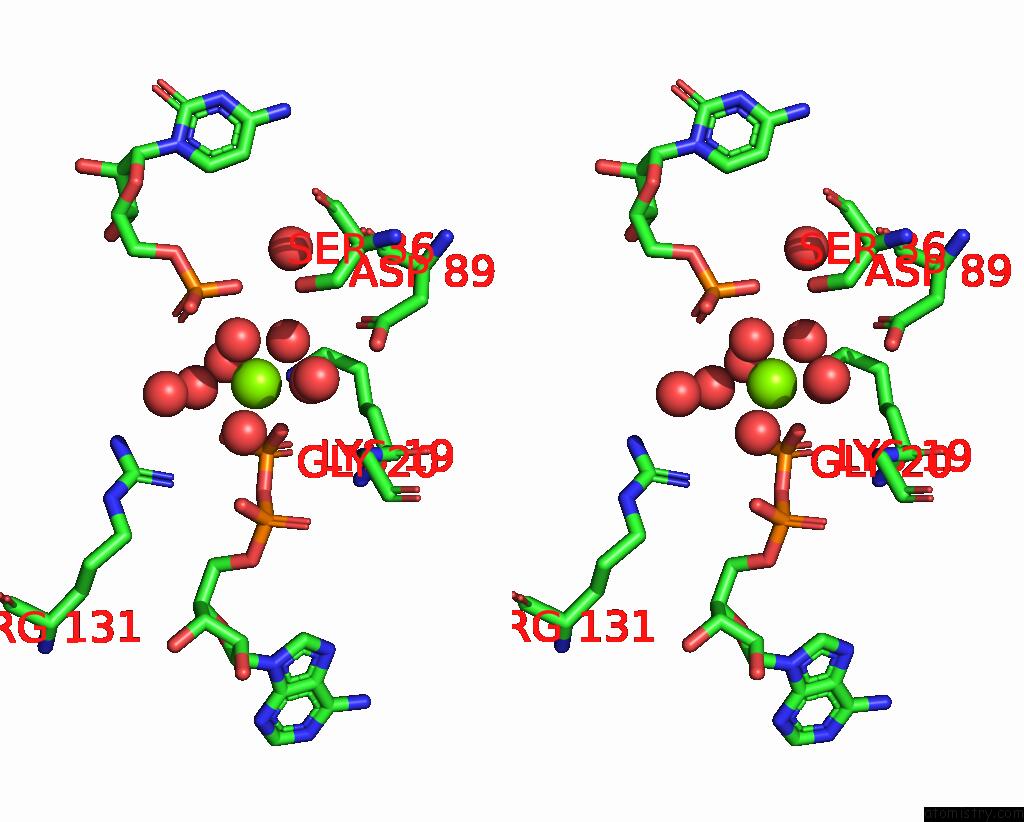
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Ump/Cmp Kinase From Slime Mold Complexed with Adp, Cmp within 5.0Å range:
|
Reference:
I.Schlichting,
J.Reinstein.
Structures of Active Conformations of Ump Kinase From Dictyostelium Discoideum Suggest Phosphoryl Transfer Is Associative. Biochemistry V. 36 9290 1997.
ISSN: ISSN 0006-2960
PubMed: 9280438
DOI: 10.1021/BI970974C
Page generated: Wed Aug 14 03:25:16 2024
ISSN: ISSN 0006-2960
PubMed: 9280438
DOI: 10.1021/BI970974C
Last articles
Cl in 5WIICl in 5WIU
Cl in 5WGZ
Cl in 5WGY
Cl in 5WHH
Cl in 5WGX
Cl in 5WGW
Cl in 5WGI
Cl in 5WGV
Cl in 5WGU