Magnesium »
PDB 3fzn-3g8c »
3g3n »
Magnesium in PDB 3g3n: PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One
Enzymatic activity of PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One
All present enzymatic activity of PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One:
3.1.4.17;
3.1.4.17;
Protein crystallography data
The structure of PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One, PDB code: 3g3n
was solved by
T.Castano,
H.Wang,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 30.00 / 2.40 |
| Space group | P 31 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 115.447, 115.447, 64.389, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 21 / 24.5 |
Other elements in 3g3n:
The structure of PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One also contains other interesting chemical elements:
| Fluorine | (F) | 2 atoms |
| Zinc | (Zn) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One
(pdb code 3g3n). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One, PDB code: 3g3n:
In total only one binding site of Magnesium was determined in the PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One, PDB code: 3g3n:
Magnesium binding site 1 out of 1 in 3g3n
Go back to
Magnesium binding site 1 out
of 1 in the PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One
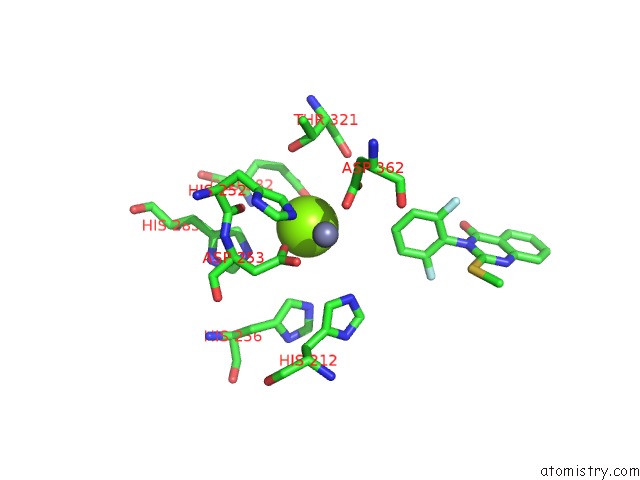
Mono view
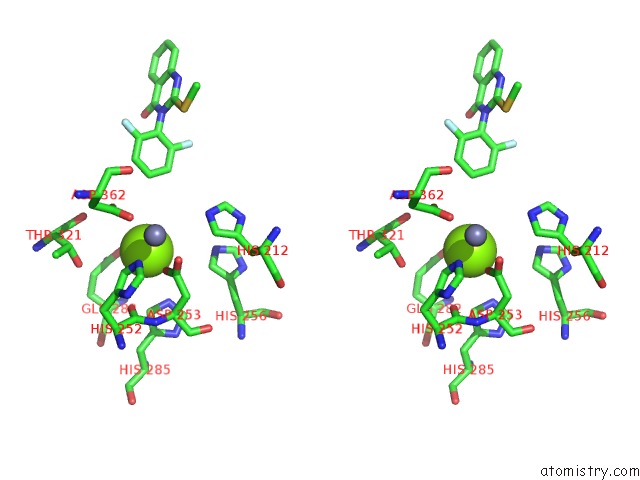
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of PDE7A Catalytic Domain in Complex with 3-(2,6- Difluorophenyl)-2-(Methylthio)Quinazolin-4(3H)-One within 5.0Å range:
|
Reference:
T.Castano,
H.Wang,
N.E.Campillo,
S.Ballester,
C.Gonzalez-Garcia,
J.Hernandez,
C.Perez,
J.Cuenca,
A.Perez-Castillo,
A.Martinez,
O.Huertas,
J.L.Gelpi,
F.J.Luque,
H.Ke,
C.Gil.
Synthesis, Structural Analysis, and Biological Evaluation of Thioxoquinazoline Derivatives As Phosphodiesterase 7 Inhibitors Chemmedchem V. 4 866 2009.
ISSN: ISSN 1860-7179
PubMed: 19350606
DOI: 10.1002/CMDC.200900043
Page generated: Sun Aug 10 21:09:59 2025
ISSN: ISSN 1860-7179
PubMed: 19350606
DOI: 10.1002/CMDC.200900043
Last articles
Mg in 7EOPMg in 7EPU
Mg in 7EOO
Mg in 7EON
Mg in 7EPK
Mg in 7EOL
Mg in 7EOK
Mg in 7EOJ
Mg in 7EOM
Mg in 7EOH