Magnesium »
PDB 3mv1-3n8b »
3my9 »
Magnesium in PDB 3my9: Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans
Protein crystallography data
The structure of Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans, PDB code: 3my9
was solved by
C.E.Quartararo,
U.Ramagopal,
J.B.Bonanno,
M.Rutter,
K.T.Bain,
S.Miller,
R.Toro,
J.M.Sauder,
S.K.Burley,
S.C.Almo,
New York Sgx Research Centerfor Structural Genomics (Nysgxrc),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 2.20 |
| Space group | I 4 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 127.993, 127.993, 98.036, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 20.2 / 25.4 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans
(pdb code 3my9). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans, PDB code: 3my9:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans, PDB code: 3my9:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 3my9
Go back to
Magnesium binding site 1 out
of 2 in the Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans
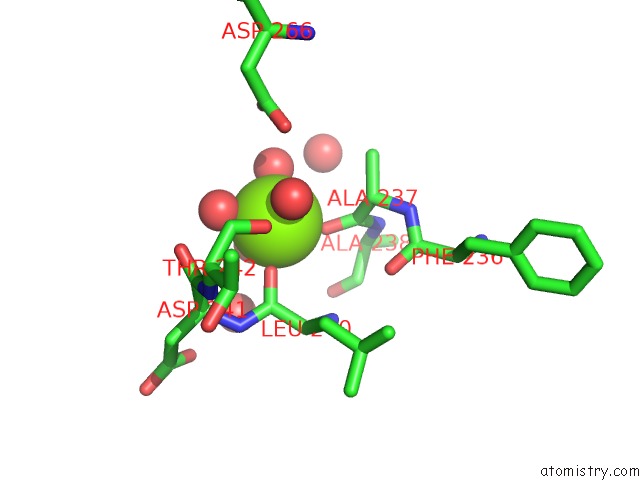
Mono view
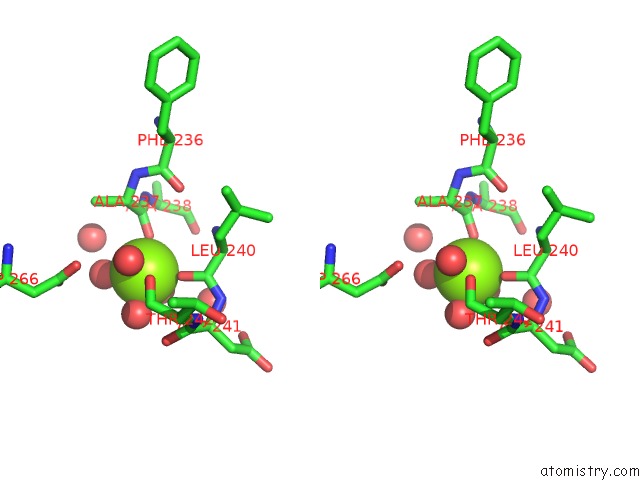
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 3my9
Go back to
Magnesium binding site 2 out
of 2 in the Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans
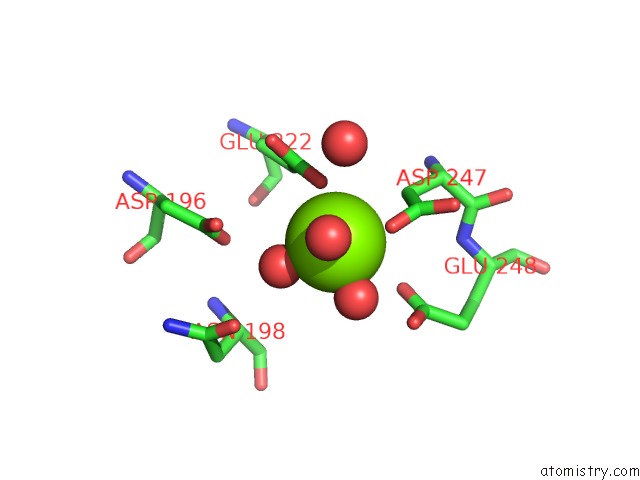
Mono view
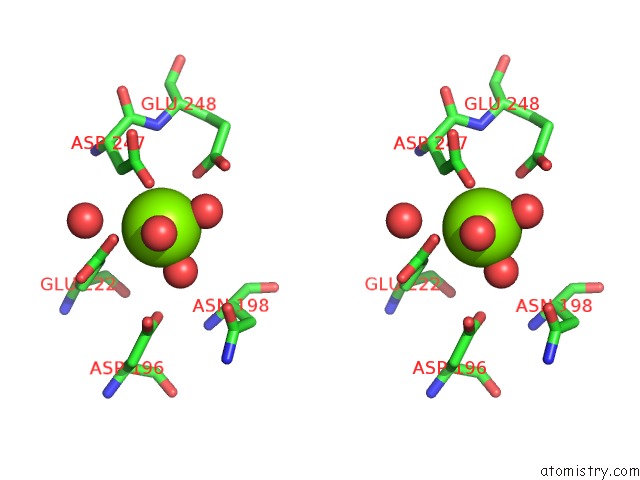
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans within 5.0Å range:
|
Reference:
C.E.Quartararo,
U.Ramagopal,
J.B.Bonanno,
M.Rutter,
K.T.Bain,
S.Miller,
R.Toro,
J.M.Sauder,
S.K.Burley,
S.C.Almo.
Crystal Structure of A Muconate Cycloisomerase From Azorhizobium Caulinodans To Be Published.
Page generated: Wed Aug 14 19:31:40 2024
Last articles
F in 4MK0F in 4MIG
F in 4MH0
F in 4MDD
F in 4MDB
F in 4MC9
F in 4MDA
F in 4MC6
F in 4MC2
F in 4MC1