Magnesium »
PDB 3mv1-3n8b »
3myr »
Magnesium in PDB 3myr: Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
Enzymatic activity of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
All present enzymatic activity of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State:
1.12.99.6;
1.12.99.6;
Protein crystallography data
The structure of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State, PDB code: 3myr
was solved by
H.Ogata,
P.Kellers,
W.Lubitz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 30.00 / 2.10 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 205.750, 216.960, 119.800, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 13.8 / 16.8 |
Other elements in 3myr:
The structure of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State also contains other interesting chemical elements:
| Nickel | (Ni) | 4 atoms |
| Iron | (Fe) | 52 atoms |
| Chlorine | (Cl) | 4 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
(pdb code 3myr). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 4 binding sites of Magnesium where determined in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State, PDB code: 3myr:
Jump to Magnesium binding site number: 1; 2; 3; 4;
In total 4 binding sites of Magnesium where determined in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State, PDB code: 3myr:
Jump to Magnesium binding site number: 1; 2; 3; 4;
Magnesium binding site 1 out of 4 in 3myr
Go back to
Magnesium binding site 1 out
of 4 in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
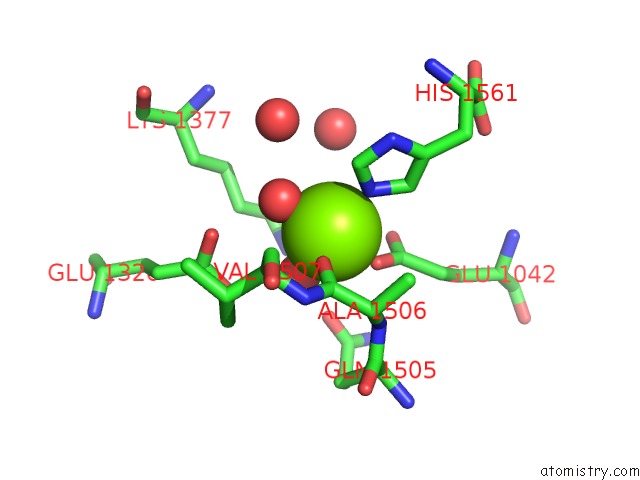
Mono view
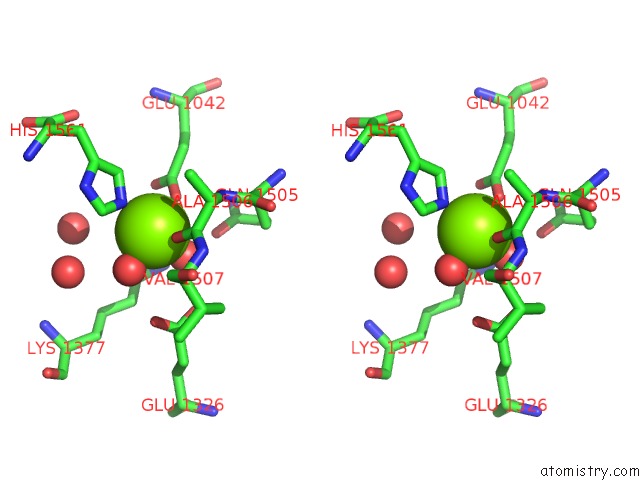
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State within 5.0Å range:
|
Magnesium binding site 2 out of 4 in 3myr
Go back to
Magnesium binding site 2 out
of 4 in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
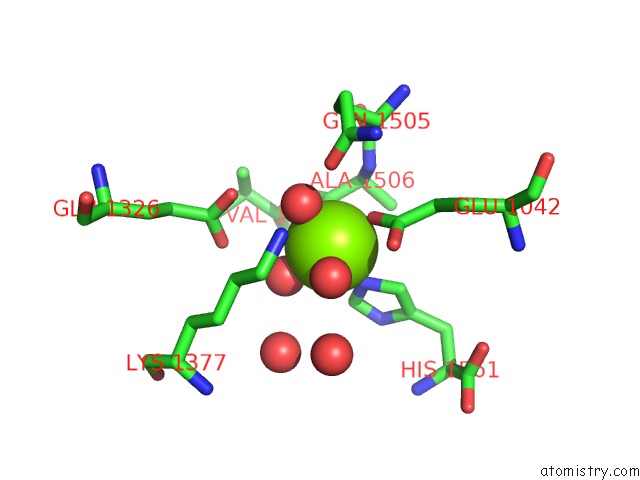
Mono view
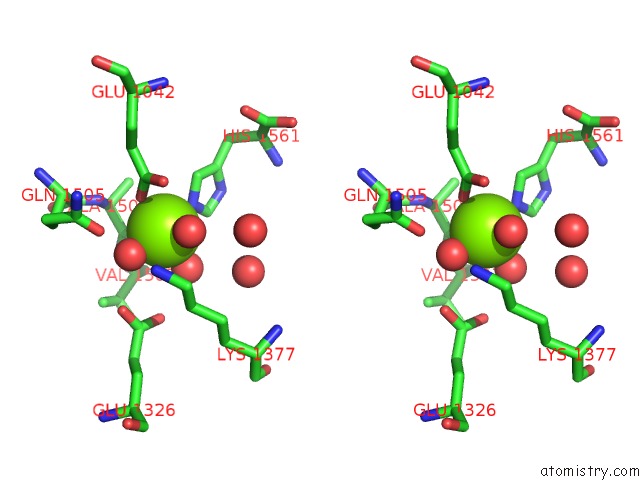
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State within 5.0Å range:
|
Magnesium binding site 3 out of 4 in 3myr
Go back to
Magnesium binding site 3 out
of 4 in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
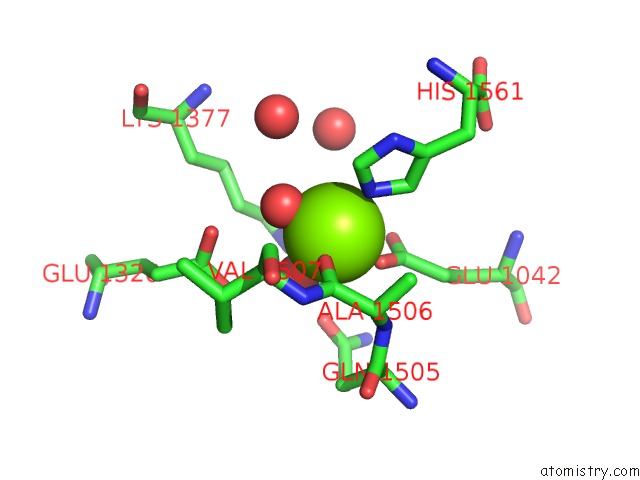
Mono view
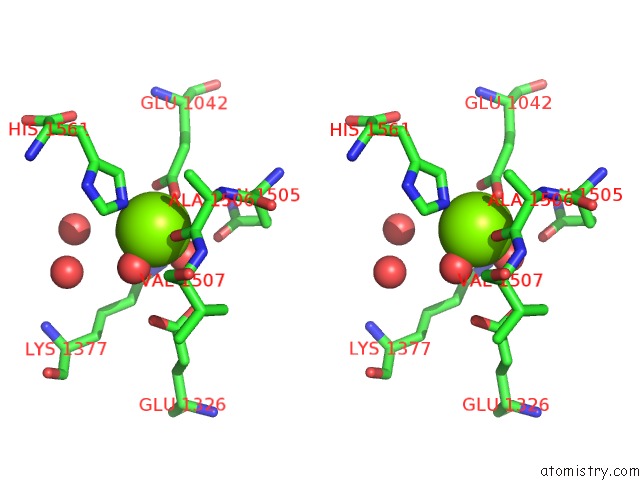
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 3 of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State within 5.0Å range:
|
Magnesium binding site 4 out of 4 in 3myr
Go back to
Magnesium binding site 4 out
of 4 in the Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State
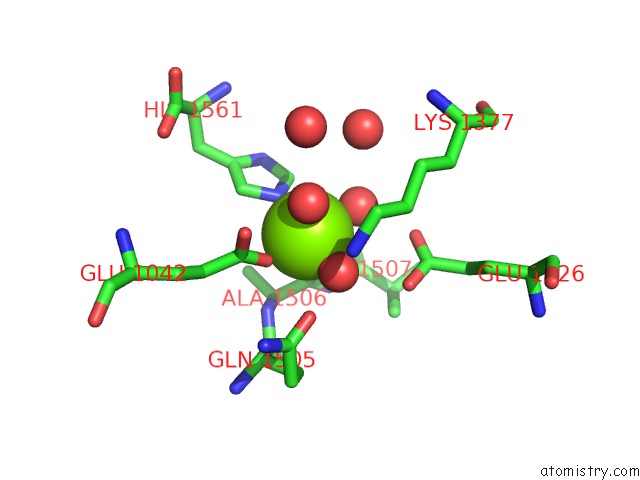
Mono view
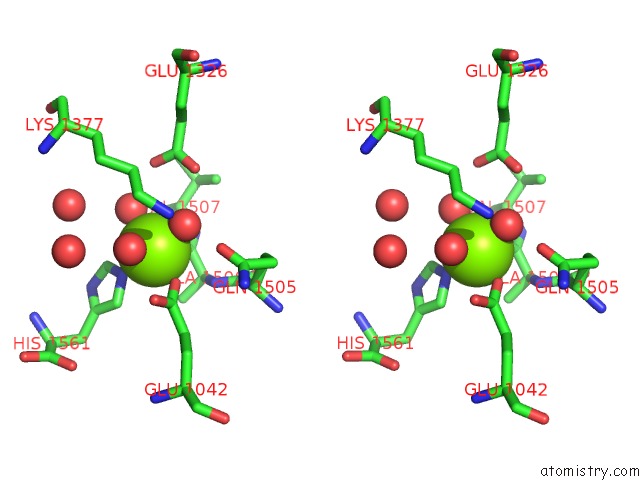
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 4 of Crystal Structure of [Nife] Hydrogenase From Allochromatium Vinosum in Its Ni-A State within 5.0Å range:
|
Reference:
H.Ogata,
P.Kellers,
W.Lubitz.
The Crystal Structure of the [Nife] Hydrogenase From the Photosynthetic Bacterium Allochromatium Vinosum: Characterization of the Oxidized Enzyme (Ni-A State). J.Mol.Biol. V. 402 428 2010.
ISSN: ISSN 0022-2836
PubMed: 20673834
DOI: 10.1016/J.JMB.2010.07.041
Page generated: Wed Aug 14 19:32:21 2024
ISSN: ISSN 0022-2836
PubMed: 20673834
DOI: 10.1016/J.JMB.2010.07.041
Last articles
Ca in 5SZMCa in 5SZL
Ca in 5SY1
Ca in 5SWI
Ca in 5SVE
Ca in 5SSX
Ca in 5SV0
Ca in 5STD
Ca in 5SSZ
Ca in 5SSY