Magnesium »
PDB 3tav-3tnf »
3tgg »
Magnesium in PDB 3tgg: A Novel Series of Potent and Selective PDE5 INHIBITOR2
Enzymatic activity of A Novel Series of Potent and Selective PDE5 INHIBITOR2
All present enzymatic activity of A Novel Series of Potent and Selective PDE5 INHIBITOR2:
3.1.4.35;
3.1.4.35;
Protein crystallography data
The structure of A Novel Series of Potent and Selective PDE5 INHIBITOR2, PDB code: 3tgg
was solved by
S.Han,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 31.00 / 1.91 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 54.493, 76.508, 76.811, 90.00, 102.42, 90.00 |
| R / Rfree (%) | 18.6 / 22.9 |
Other elements in 3tgg:
The structure of A Novel Series of Potent and Selective PDE5 INHIBITOR2 also contains other interesting chemical elements:
| Zinc | (Zn) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the A Novel Series of Potent and Selective PDE5 INHIBITOR2
(pdb code 3tgg). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the A Novel Series of Potent and Selective PDE5 INHIBITOR2, PDB code: 3tgg:
In total only one binding site of Magnesium was determined in the A Novel Series of Potent and Selective PDE5 INHIBITOR2, PDB code: 3tgg:
Magnesium binding site 1 out of 1 in 3tgg
Go back to
Magnesium binding site 1 out
of 1 in the A Novel Series of Potent and Selective PDE5 INHIBITOR2
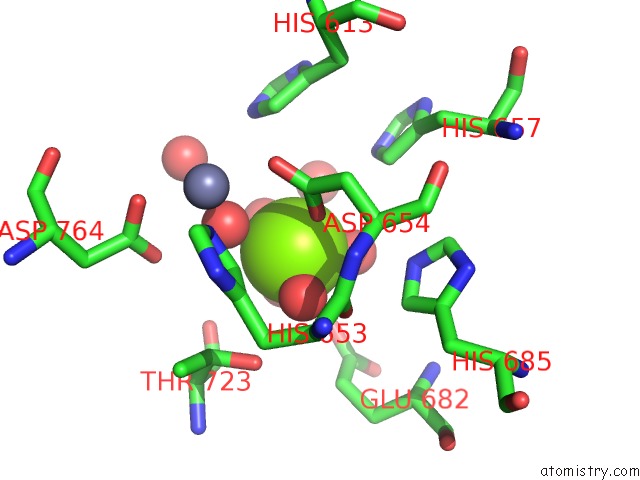
Mono view
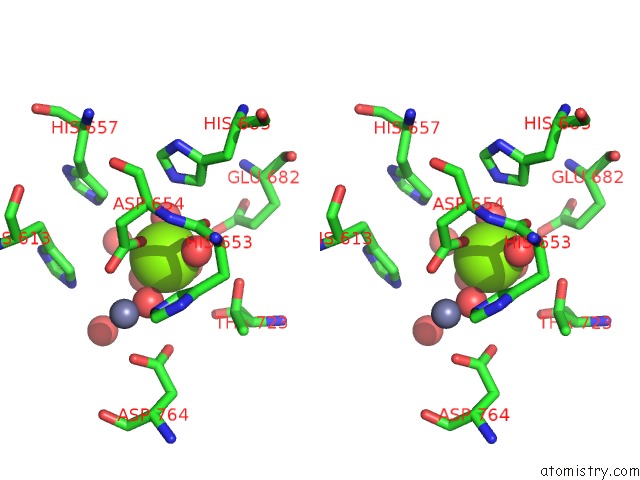
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of A Novel Series of Potent and Selective PDE5 INHIBITOR2 within 5.0Å range:
|
Reference:
R.O.Hughes,
T.Maddux,
D.Joseph Rogier,
S.Lu,
J.K.Walker,
E.Jon Jacobsen,
J.M.Rumsey,
Y.Zheng,
A.Macinnes,
B.R.Bond,
S.Han.
Investigation of the Pyrazinones As PDE5 Inhibitors: Evaluation of Regioisomeric Projections Into the Solvent Region. Bioorg.Med.Chem.Lett. V. 21 6348 2011.
ISSN: ISSN 0960-894X
PubMed: 21955943
DOI: 10.1016/J.BMCL.2011.08.106
Page generated: Thu Aug 15 12:06:13 2024
ISSN: ISSN 0960-894X
PubMed: 21955943
DOI: 10.1016/J.BMCL.2011.08.106
Last articles
Ca in 5VOLCa in 5VRX
Ca in 5VRW
Ca in 5VR0
Ca in 5VPL
Ca in 5VPH
Ca in 5VMS
Ca in 5VPG
Ca in 5VOF
Ca in 5VMA