Magnesium »
PDB 3u8x-3ump »
3ucd »
Magnesium in PDB 3ucd: Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
Enzymatic activity of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
All present enzymatic activity of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep:
4.2.1.11;
4.2.1.11;
Protein crystallography data
The structure of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep, PDB code: 3ucd
was solved by
J.Qin,
G.Chai,
J.Brewer,
L.Lovelace,
L.Lebioda,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 44.58 / 1.41 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 105.708, 118.623, 67.579, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16.1 / 19.8 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
(pdb code 3ucd). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 4 binding sites of Magnesium where determined in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep, PDB code: 3ucd:
Jump to Magnesium binding site number: 1; 2; 3; 4;
In total 4 binding sites of Magnesium where determined in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep, PDB code: 3ucd:
Jump to Magnesium binding site number: 1; 2; 3; 4;
Magnesium binding site 1 out of 4 in 3ucd
Go back to
Magnesium binding site 1 out
of 4 in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
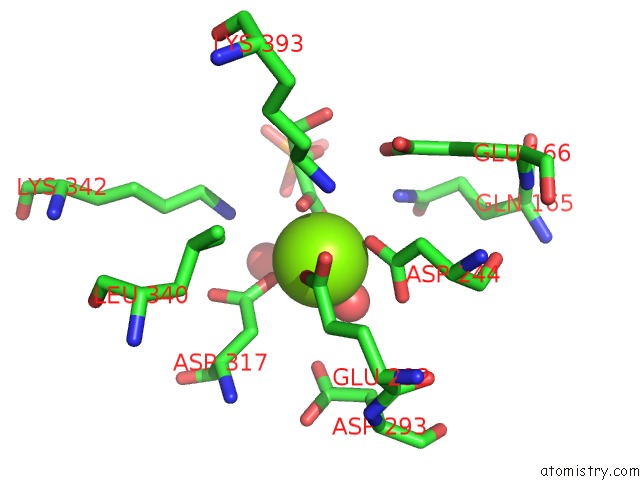
Mono view
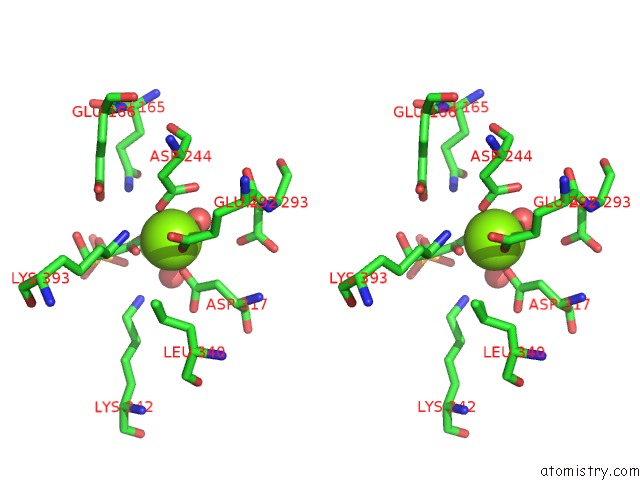
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep within 5.0Å range:
|
Magnesium binding site 2 out of 4 in 3ucd
Go back to
Magnesium binding site 2 out
of 4 in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
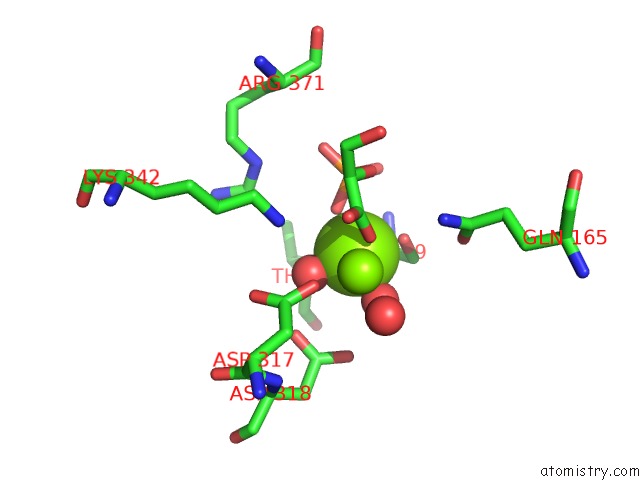
Mono view
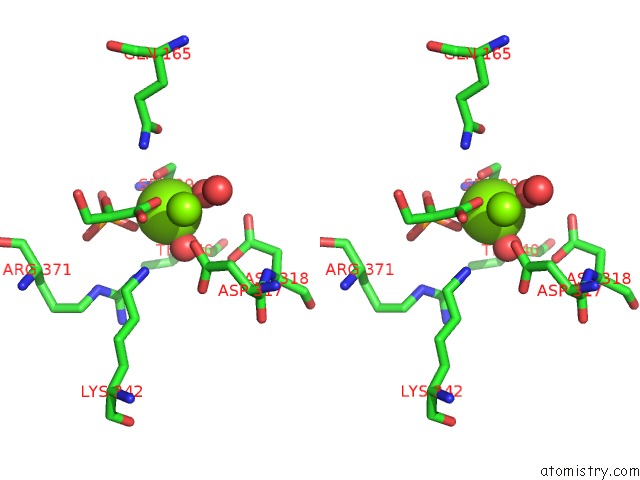
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep within 5.0Å range:
|
Magnesium binding site 3 out of 4 in 3ucd
Go back to
Magnesium binding site 3 out
of 4 in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
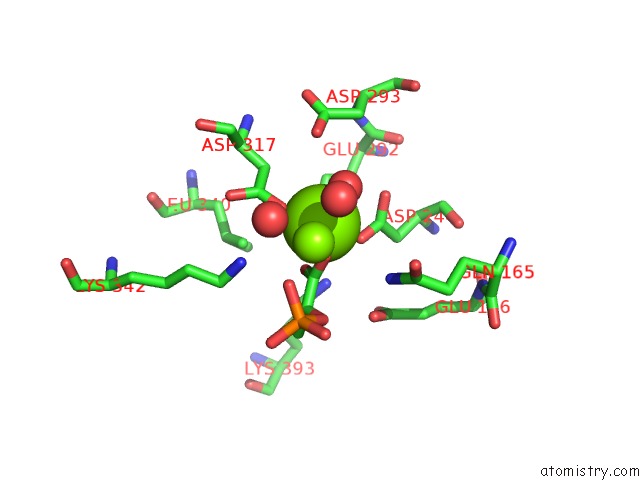
Mono view
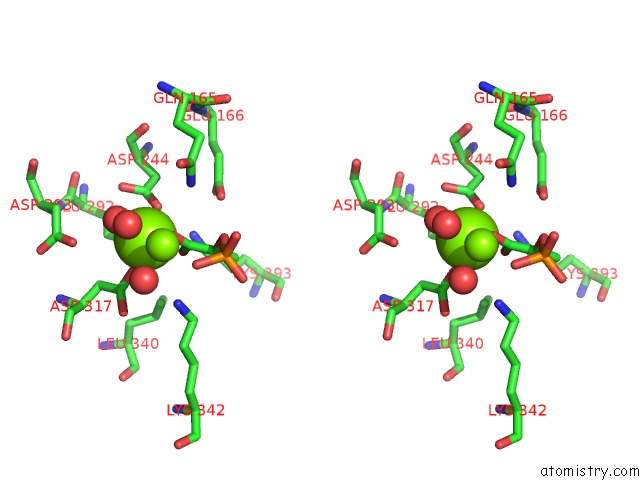
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 3 of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep within 5.0Å range:
|
Magnesium binding site 4 out of 4 in 3ucd
Go back to
Magnesium binding site 4 out
of 4 in the Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep
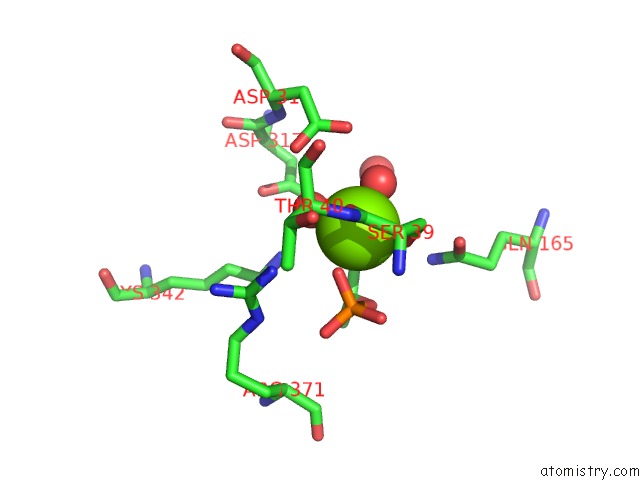
Mono view
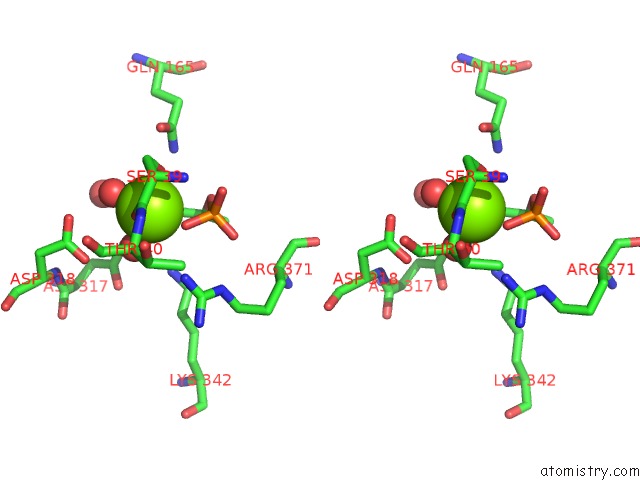
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 4 of Asymmetric Complex of Human Neuron Specific Enolase-2-Pga/Pep within 5.0Å range:
|
Reference:
J.Qin,
G.Chai,
J.M.Brewer,
L.L.Lovelace,
L.Lebioda.
Structures of Asymmetric Complexes of Human Neuron Specific Enolase with Resolved Substrate and Product and An Analogous Complex with Two Inhibitors Indicate Subunit Interaction and Inhibitor Cooperativity. J.Inorg.Biochem. V. 111 187 2012.
ISSN: ISSN 0162-0134
PubMed: 22437160
DOI: 10.1016/J.JINORGBIO.2012.02.011
Page generated: Thu Aug 15 12:27:14 2024
ISSN: ISSN 0162-0134
PubMed: 22437160
DOI: 10.1016/J.JINORGBIO.2012.02.011
Last articles
F in 4HU1F in 4HVS
F in 4HW7
F in 4HUA
F in 4HU9
F in 4HQJ
F in 4HT3
F in 4HLQ
F in 4HT0
F in 4HNA