Magnesium »
PDB 4cgl-4cs4 »
4crq »
Magnesium in PDB 4crq: Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S
Enzymatic activity of Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S
All present enzymatic activity of Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S:
3.2.1.39;
3.2.1.39;
Protein crystallography data
The structure of Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S, PDB code: 4crq
was solved by
A.Labourel,
M.Jam,
L.Legentil,
B.Sylla,
E.Ficko-Blean,
J.H.Hehemann,
V.Ferrieres,
M.Czjzek,
G.Michel,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 48.98 / 1.50 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 67.108, 68.061, 143.282, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 14.802 / 18.576 |
Other elements in 4crq:
The structure of Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S also contains other interesting chemical elements:
| Chlorine | (Cl) | 3 atoms |
| Calcium | (Ca) | 2 atoms |
| Sodium | (Na) | 2 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S
(pdb code 4crq). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S, PDB code: 4crq:
In total only one binding site of Magnesium was determined in the Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S, PDB code: 4crq:
Magnesium binding site 1 out of 1 in 4crq
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S
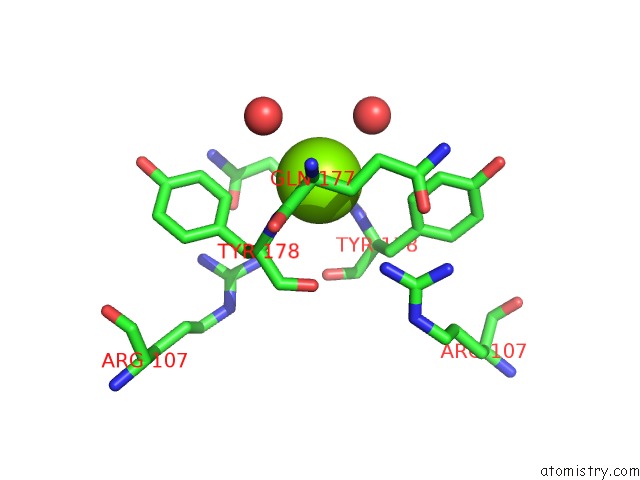
Mono view
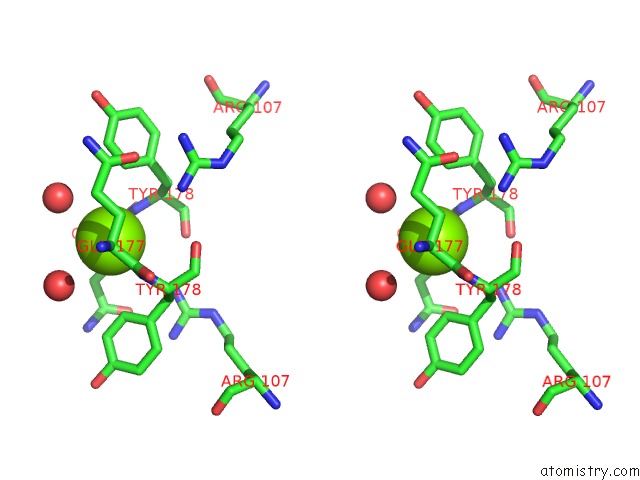
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of the Catalytic Domain of the Modular Laminarinase Zglamc Mutant E142S within 5.0Å range:
|
Reference:
A.Labourel,
M.Jam,
L.Legentil,
B.Sylla,
J.H.Hehemann,
V.Ferrieres,
M.Czjzek,
G.Michel.
Structural and Biochemical Characterization of the Laminarina Zglamc[GH16] From Zobellia Galactanivorans Suggests Preferred Recognition of Branched Laminarin Acta Crystallogr.,Sect.D V. 71 173 2015.
ISSN: ISSN 0907-4449
DOI: 10.1107/S139900471402450X
Page generated: Mon Aug 11 07:18:30 2025
ISSN: ISSN 0907-4449
DOI: 10.1107/S139900471402450X
Last articles
Mg in 4DUZMg in 4DUY
Mg in 4DR7
Mg in 4DR6
Mg in 4DR5
Mg in 4DUX
Mg in 4DUW
Mg in 4DUV
Mg in 4DUO
Mg in 4DUG