Magnesium »
PDB 4ibd-4iir »
4ibd »
Magnesium in PDB 4ibd: Ebola Virus VP35 Bound to Small Molecule
Protein crystallography data
The structure of Ebola Virus VP35 Bound to Small Molecule, PDB code: 4ibd
was solved by
C.S.Brown,
D.W.Leung,
W.Xu,
D.M.Borek,
Z.Otwinowski,
P.Ramanan,
A.J.Stubbs,
D.S.Peterson,
J.M.Binning,
G.K.Amarasinghe,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 35.38 / 1.84 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 51.562, 65.485, 72.596, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.7 / 23.3 |
Other elements in 4ibd:
The structure of Ebola Virus VP35 Bound to Small Molecule also contains other interesting chemical elements:
| Bromine | (Br) | 2 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Ebola Virus VP35 Bound to Small Molecule
(pdb code 4ibd). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Ebola Virus VP35 Bound to Small Molecule, PDB code: 4ibd:
In total only one binding site of Magnesium was determined in the Ebola Virus VP35 Bound to Small Molecule, PDB code: 4ibd:
Magnesium binding site 1 out of 1 in 4ibd
Go back to
Magnesium binding site 1 out
of 1 in the Ebola Virus VP35 Bound to Small Molecule
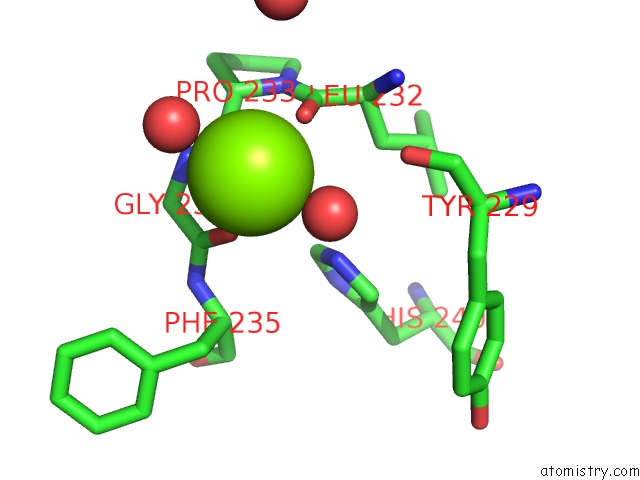
Mono view
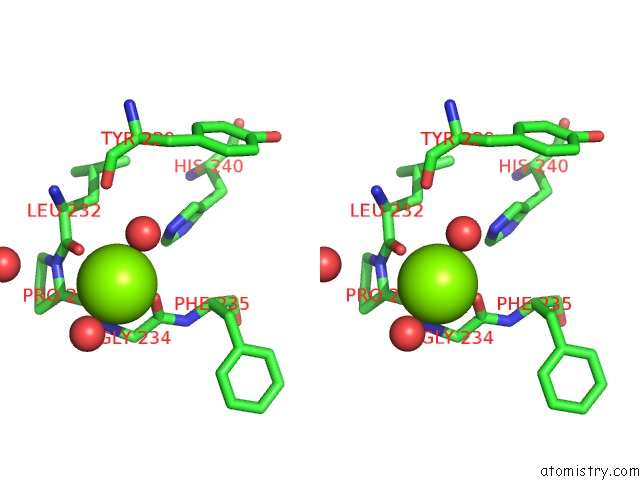
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Ebola Virus VP35 Bound to Small Molecule within 5.0Å range:
|
Reference:
C.S.Brown,
M.S.Lee,
D.W.Leung,
T.Wang,
W.Xu,
P.Luthra,
M.Anantpadma,
R.S.Shabman,
L.M.Melito,
K.S.Macmillan,
D.M.Borek,
Z.Otwinowski,
P.Ramanan,
A.J.Stubbs,
D.S.Peterson,
J.M.Binning,
M.Tonelli,
M.A.Olson,
R.A.Davey,
J.M.Ready,
C.F.Basler,
G.K.Amarasinghe.
In Silico Derived Small Molecules Bind the Filovirus VP35 Protein and Inhibit Its Polymerase Cofactor Activity. J.Mol.Biol. V. 426 2045 2014.
ISSN: ISSN 0022-2836
PubMed: 24495995
DOI: 10.1016/J.JMB.2014.01.010
Page generated: Fri Aug 16 16:38:13 2024
ISSN: ISSN 0022-2836
PubMed: 24495995
DOI: 10.1016/J.JMB.2014.01.010
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF