Magnesium »
PDB 5cr7-5d2j »
5d1f »
Magnesium in PDB 5d1f: Crystal Structure of Maize Pdrp Bound with Amp and HG2+
Enzymatic activity of Crystal Structure of Maize Pdrp Bound with Amp and HG2+
All present enzymatic activity of Crystal Structure of Maize Pdrp Bound with Amp and HG2+:
2.7.11.32; 2.7.4.27;
2.7.11.32; 2.7.4.27;
Protein crystallography data
The structure of Crystal Structure of Maize Pdrp Bound with Amp and HG2+, PDB code: 5d1f
was solved by
L.Jiang,
Z.Chen,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 48.68 / 3.40 |
| Space group | P 4 3 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 175.514, 175.514, 175.514, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 22.3 / 24.3 |
Other elements in 5d1f:
The structure of Crystal Structure of Maize Pdrp Bound with Amp and HG2+ also contains other interesting chemical elements:
| Mercury | (Hg) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of Maize Pdrp Bound with Amp and HG2+
(pdb code 5d1f). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of Maize Pdrp Bound with Amp and HG2+, PDB code: 5d1f:
In total only one binding site of Magnesium was determined in the Crystal Structure of Maize Pdrp Bound with Amp and HG2+, PDB code: 5d1f:
Magnesium binding site 1 out of 1 in 5d1f
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of Maize Pdrp Bound with Amp and HG2+
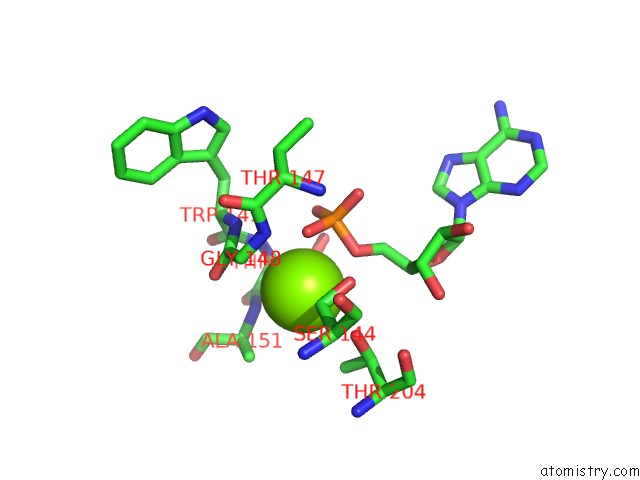
Mono view
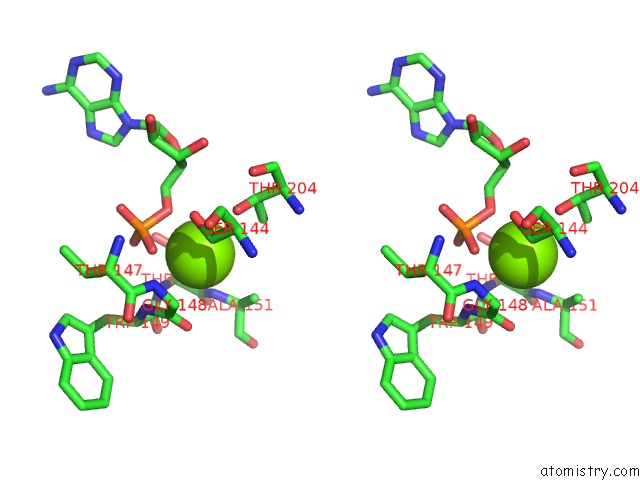
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of Maize Pdrp Bound with Amp and HG2+ within 5.0Å range:
|
Reference:
L.Jiang,
Y.B.Chen,
J.Zheng,
Z.Chen,
Y.Liu,
Y.Tao,
W.Wu,
Z.Chen,
B.C.Wang.
Structural Basis of Reversible Phosphorylation By Maize Pyruvate Orthophosphate Dikinase Regulatory Protein Plant Physiol. V. 170 732 2016.
ISSN: ESSN 1532-2548
PubMed: 26620526
DOI: 10.1104/PP.15.01709
Page generated: Sun Sep 29 02:26:31 2024
ISSN: ESSN 1532-2548
PubMed: 26620526
DOI: 10.1104/PP.15.01709
Last articles
Cl in 5WDJCl in 5WE8
Cl in 5WBG
Cl in 5WDW
Cl in 5WE7
Cl in 5WBV
Cl in 5WDC
Cl in 5WCT
Cl in 5WCI
Cl in 5WBQ