Magnesium »
PDB 5d2k-5dar »
5d9b »
Magnesium in PDB 5d9b: Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data)
Protein crystallography data
The structure of Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data), PDB code: 5d9b
was solved by
K.Yamashita,
D.Pan,
T.Okuda,
T.Murai,
A.Kodan,
T.Yamaguchi,
K.Gomi,
N.Kajiyama,
H.Kato,
H.Ago,
M.Yamamoto,
T.Nakatsu,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 10.00 / 1.50 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 48.210, 77.590, 84.760, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.5 / 23.2 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data)
(pdb code 5d9b). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data), PDB code: 5d9b:
In total only one binding site of Magnesium was determined in the Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data), PDB code: 5d9b:
Magnesium binding site 1 out of 1 in 5d9b
Go back to
Magnesium binding site 1 out
of 1 in the Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data)
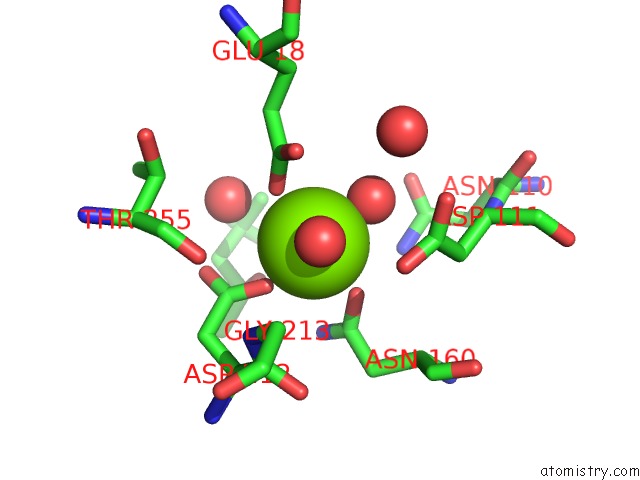
Mono view
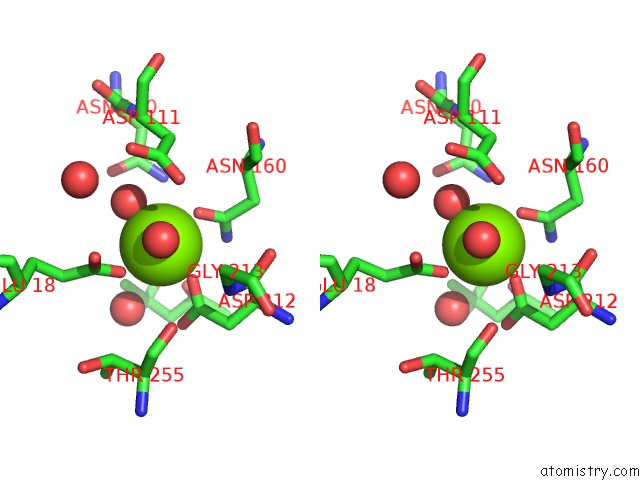
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Luciferin-Regenerating Enzyme Solved By Siras Using Xfel (Refined Against Native Data) within 5.0Å range:
|
Reference:
K.Yamashita,
D.Pan,
T.Okuda,
M.Sugahara,
A.Kodan,
T.Yamaguchi,
T.Murai,
K.Gomi,
N.Kajiyama,
E.Mizohata,
M.Suzuki,
E.Nango,
K.Tono,
Y.Joti,
T.Kameshima,
J.Park,
C.Song,
T.Hatsui,
M.Yabashi,
S.Iwata,
H.Kato,
H.Ago,
M.Yamamoto,
T.Nakatsu.
An Isomorphous Replacement Method For Efficient De Novo Phasing For Serial Femtosecond Crystallography. Sci Rep V. 5 14017 2015.
ISSN: ESSN 2045-2322
PubMed: 26360462
DOI: 10.1038/SREP14017
Page generated: Tue Aug 12 06:50:53 2025
ISSN: ESSN 2045-2322
PubMed: 26360462
DOI: 10.1038/SREP14017
Last articles
Mg in 5HKVMg in 5HR6
Mg in 5HQL
Mg in 5HQW
Mg in 5HQM
Mg in 5HQC
Mg in 5HQB
Mg in 5HQ8
Mg in 5HQA
Mg in 5HQ4