Magnesium »
PDB 5egy-5etr »
5erv »
Magnesium in PDB 5erv: Ternary Complex of Gephe - Adp - Tungsten Cluster
Enzymatic activity of Ternary Complex of Gephe - Adp - Tungsten Cluster
All present enzymatic activity of Ternary Complex of Gephe - Adp - Tungsten Cluster:
2.10.1.1; 2.7.7.75;
2.10.1.1; 2.7.7.75;
Protein crystallography data
The structure of Ternary Complex of Gephe - Adp - Tungsten Cluster, PDB code: 5erv
was solved by
V.B.Kasaragod,
H.Schindelin,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 65.83 / 1.80 |
| Space group | I 2 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 87.661, 99.675, 113.237, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 15.6 / 20.3 |
Other elements in 5erv:
The structure of Ternary Complex of Gephe - Adp - Tungsten Cluster also contains other interesting chemical elements:
| Tungsten | (W) | 10 atoms |
| Calcium | (Ca) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Ternary Complex of Gephe - Adp - Tungsten Cluster
(pdb code 5erv). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Ternary Complex of Gephe - Adp - Tungsten Cluster, PDB code: 5erv:
In total only one binding site of Magnesium was determined in the Ternary Complex of Gephe - Adp - Tungsten Cluster, PDB code: 5erv:
Magnesium binding site 1 out of 1 in 5erv
Go back to
Magnesium binding site 1 out
of 1 in the Ternary Complex of Gephe - Adp - Tungsten Cluster
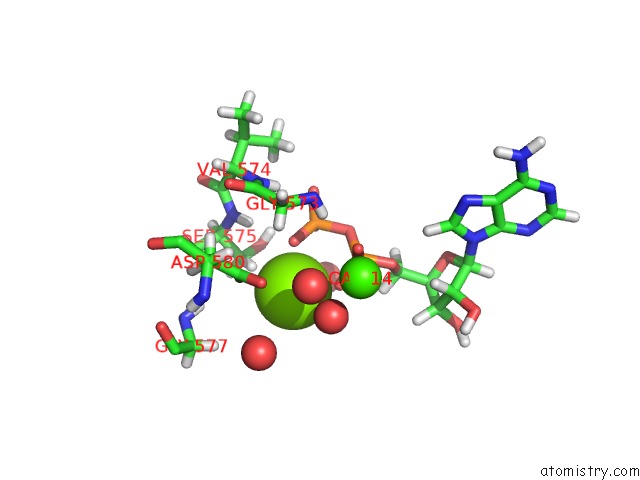
Mono view
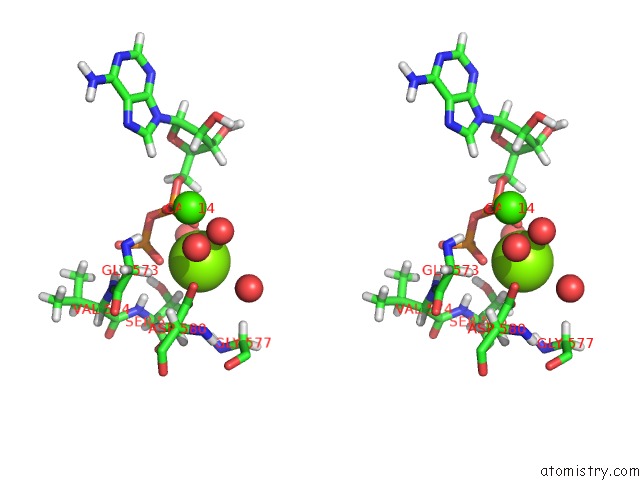
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Ternary Complex of Gephe - Adp - Tungsten Cluster within 5.0Å range:
|
Reference:
V.B.Kasaragod,
H.Schindelin.
Structural Framework For Metal Incorporation During Molybdenum Cofactor Biosynthesis. Structure V. 24 782 2016.
ISSN: ISSN 0969-2126
PubMed: 27112598
DOI: 10.1016/J.STR.2016.02.023
Page generated: Sun Sep 29 03:54:19 2024
ISSN: ISSN 0969-2126
PubMed: 27112598
DOI: 10.1016/J.STR.2016.02.023
Last articles
Cl in 7T1KCl in 7T0C
Cl in 7SZE
Cl in 7T0E
Cl in 7SZF
Cl in 7SZQ
Cl in 7SZP
Cl in 7SZH
Cl in 7SZG
Cl in 7SYY