Magnesium »
PDB 5z79-5zki »
5z9p »
Magnesium in PDB 5z9p: Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment
Enzymatic activity of Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment
All present enzymatic activity of Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment:
5.99.1.3;
5.99.1.3;
Protein crystallography data
The structure of Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment, PDB code: 5z9p
was solved by
X.Huang,
H.Zhou,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 70.40 / 1.45 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 143.103, 55.660, 51.055, 90.00, 100.27, 90.00 |
| R / Rfree (%) | 19.3 / 21 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment
(pdb code 5z9p). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment, PDB code: 5z9p:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment, PDB code: 5z9p:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 5z9p
Go back to
Magnesium binding site 1 out
of 2 in the Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment
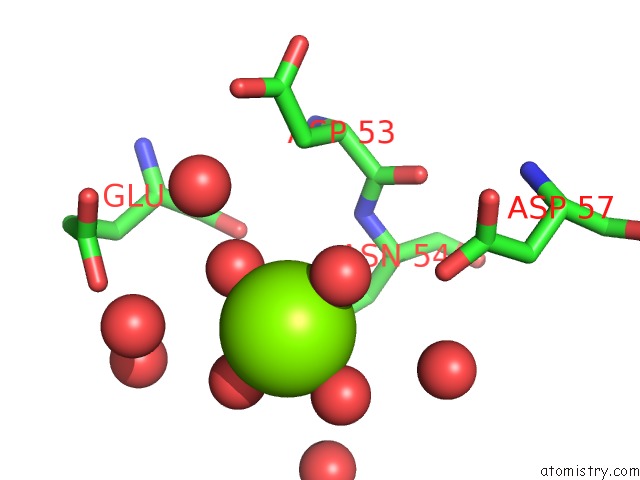
Mono view
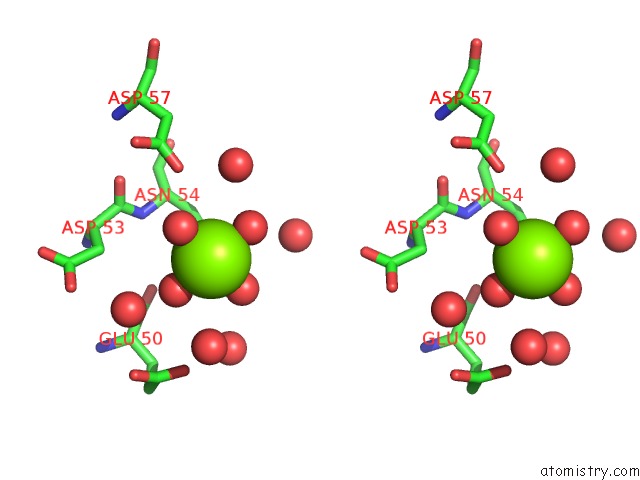
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 5z9p
Go back to
Magnesium binding site 2 out
of 2 in the Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment
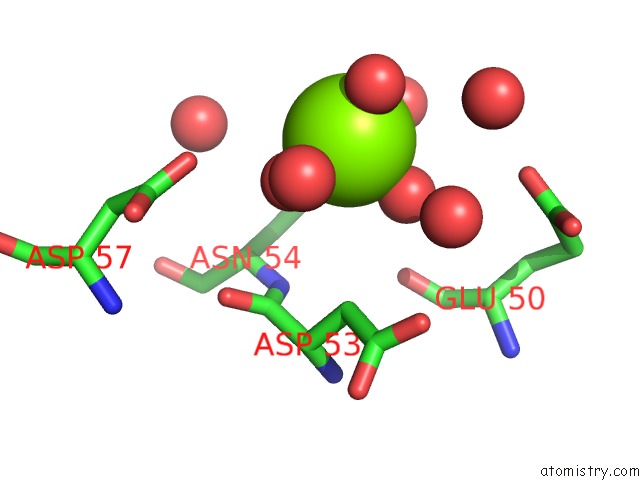
Mono view
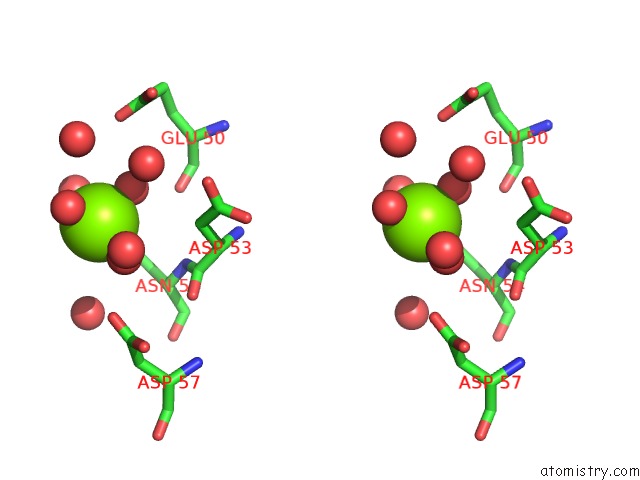
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Bacterial Gyrb Atpase Domain in Complex with A Chemical Fragment within 5.0Å range:
|
Reference:
X.Huang,
J.Guo,
Q.Liu,
Q.Gu,
J.Xu,
H.Zhou.
Identification of An Auxiliary Druggable Pocket in the Dna Gyrase Atpase Domain Using Fragment Probes Medchemcomm V. 9 1619 2018.
ISSN: ESSN 2040-2511
PubMed: 30429968
DOI: 10.1039/C8MD00148K
Page generated: Mon Sep 30 11:47:21 2024
ISSN: ESSN 2040-2511
PubMed: 30429968
DOI: 10.1039/C8MD00148K
Last articles
F in 4IVOF in 4IV4
F in 4IVM
F in 4IV2
F in 4IUI
F in 4IN4
F in 4IU7
F in 4ITI
F in 4IUE
F in 4IRU