Magnesium »
PDB 6a1m-6aad »
6a5u »
Magnesium in PDB 6a5u: Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation
Enzymatic activity of Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation
All present enzymatic activity of Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation:
2.7.7.6;
2.7.7.6;
Other elements in 6a5u:
The structure of Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation also contains other interesting chemical elements:
| Zinc | (Zn) | 8 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation
(pdb code 6a5u). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation, PDB code: 6a5u:
In total only one binding site of Magnesium was determined in the Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation, PDB code: 6a5u:
Magnesium binding site 1 out of 1 in 6a5u
Go back to
Magnesium binding site 1 out
of 1 in the Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation
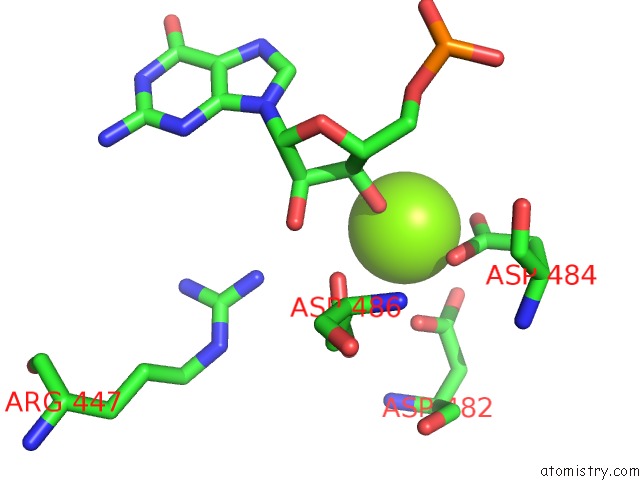
Mono view
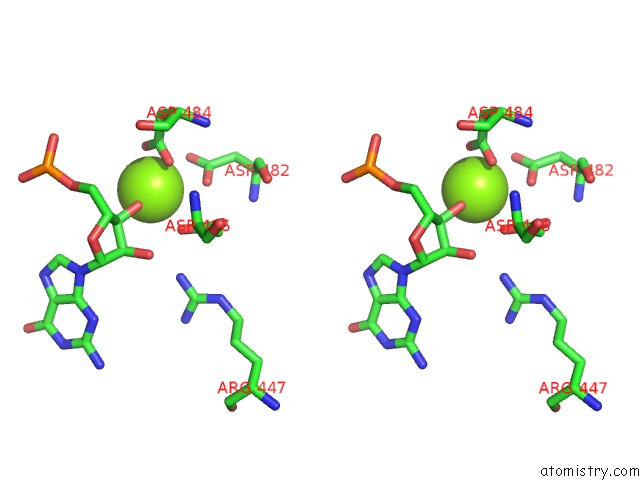
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Rna Polymerase II Elongation Complex Stalled at Shl(-1) of the Nucleosome, with Foreign Dna, Tilt Conformation within 5.0Å range:
|
Reference:
T.Kujirai,
H.Ehara,
Y.Fujino,
M.Shirouzu,
S.I.Sekine,
H.Kurumizaka.
Structural Basis of the Nucleosome Transition During Rna Polymerase II Passage. Science V. 362 595 2018.
ISSN: ESSN 1095-9203
PubMed: 30287617
DOI: 10.1126/SCIENCE.AAU9904
Page generated: Mon Sep 30 19:00:22 2024
ISSN: ESSN 1095-9203
PubMed: 30287617
DOI: 10.1126/SCIENCE.AAU9904
Last articles
Cl in 7T0CCl in 7SZE
Cl in 7T0E
Cl in 7SZF
Cl in 7SZQ
Cl in 7SZP
Cl in 7SZH
Cl in 7SZG
Cl in 7SYY
Cl in 7SXL