Magnesium »
PDB 6bcr-6bqz »
6bjs »
Magnesium in PDB 6bjs: Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin
Enzymatic activity of Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin
All present enzymatic activity of Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin:
2.7.7.6;
2.7.7.6;
Other elements in 6bjs:
The structure of Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin
(pdb code 6bjs). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin, PDB code: 6bjs:
In total only one binding site of Magnesium was determined in the Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin, PDB code: 6bjs:
Magnesium binding site 1 out of 1 in 6bjs
Go back to
Magnesium binding site 1 out
of 1 in the Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin
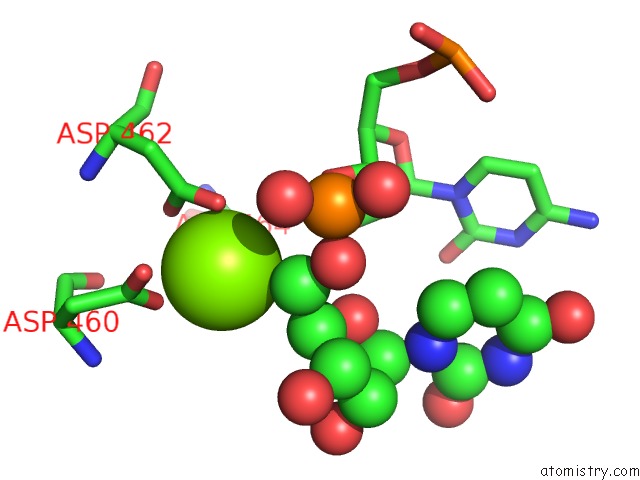
Mono view
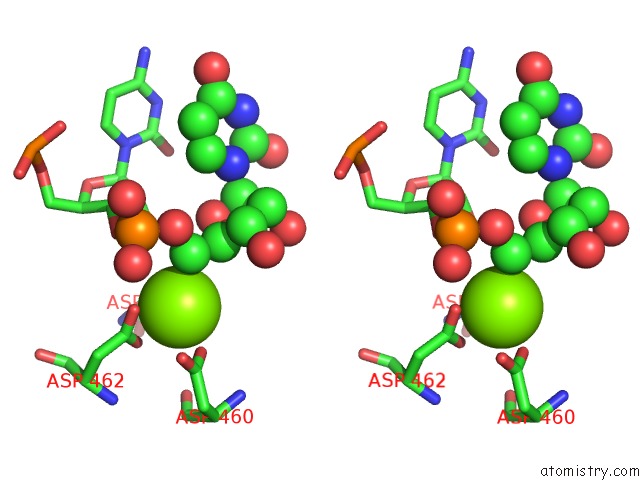
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Cryoem Structure of E.Coli His Pause Elongation Complex Without Pause Hairpin within 5.0Å range:
|
Reference:
J.Y.Kang,
T.V.Mishanina,
M.J.Bellecourt,
R.A.Mooney,
S.A.Darst,
R.Landick.
Rna Polymerase Accommodates A Pause Rna Hairpin By Global Conformational Rearrangements That Prolong Pausing. Mol. Cell V. 69 802 2018.
ISSN: ISSN 1097-4164
PubMed: 29499135
DOI: 10.1016/J.MOLCEL.2018.01.018
Page generated: Mon Sep 30 19:46:17 2024
ISSN: ISSN 1097-4164
PubMed: 29499135
DOI: 10.1016/J.MOLCEL.2018.01.018
Last articles
Zn in 9JYWZn in 9IR4
Zn in 9IR3
Zn in 9GMX
Zn in 9GMW
Zn in 9JEJ
Zn in 9ERF
Zn in 9ERE
Zn in 9EGV
Zn in 9EGW