Magnesium »
PDB 6u1y-6ufh »
6u43 »
Magnesium in PDB 6u43: Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form)
Protein crystallography data
The structure of Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form), PDB code: 6u43
was solved by
M.K.Fenwick,
S.E.Ealick,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 54.26 / 1.40 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 57.195, 62.524, 63.396, 64.21, 69.68, 80.51 |
| R / Rfree (%) | 16.5 / 19.2 |
Other elements in 6u43:
The structure of Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form) also contains other interesting chemical elements:
| Chlorine | (Cl) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form)
(pdb code 6u43). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form), PDB code: 6u43:
In total only one binding site of Magnesium was determined in the Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form), PDB code: 6u43:
Magnesium binding site 1 out of 1 in 6u43
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form)
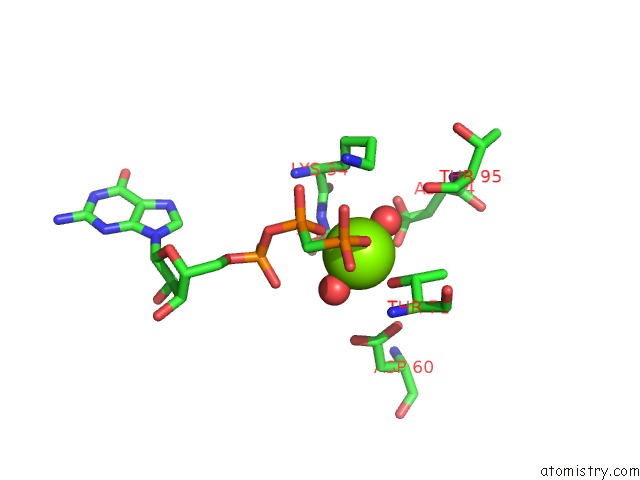
Mono view
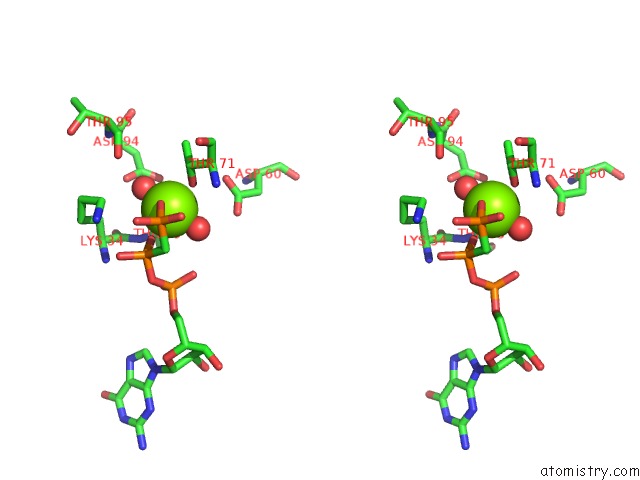
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of Methanoperedens Nitroreducens Elongation Factor 2 H595N Bound to Gmppcp and Magnesium (Triclinic Crystal Form) within 5.0Å range:
|
Reference:
M.K.Fenwick,
S.E.Ealick.
Structural Basis of Elongation Factor 2 Switching Curr Res Struct Biol V. 2 25 2020.
ISSN: ESSN 2665-928X
DOI: 10.1016/J.CRSTBI.2020.02.001
Page generated: Tue Oct 1 20:49:07 2024
ISSN: ESSN 2665-928X
DOI: 10.1016/J.CRSTBI.2020.02.001
Last articles
Ca in 2XGQCa in 2XHN
Ca in 2XGP
Ca in 2XJ7
Ca in 2XHI
Ca in 2XHJ
Ca in 2XHH
Ca in 2XF2
Ca in 2XFV
Ca in 2XFG