Magnesium »
PDB 6u1y-6ufh »
6ubz »
Magnesium in PDB 6ubz: Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5
Protein crystallography data
The structure of Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5, PDB code: 6ubz
was solved by
E.T.Yukl,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 47.35 / 1.83 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 110.325, 93.403, 187.987, 90.00, 95.20, 90.00 |
| R / Rfree (%) | 16.2 / 18.9 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5
(pdb code 6ubz). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 4 binding sites of Magnesium where determined in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5, PDB code: 6ubz:
Jump to Magnesium binding site number: 1; 2; 3; 4;
In total 4 binding sites of Magnesium where determined in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5, PDB code: 6ubz:
Jump to Magnesium binding site number: 1; 2; 3; 4;
Magnesium binding site 1 out of 4 in 6ubz
Go back to
Magnesium binding site 1 out
of 4 in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5
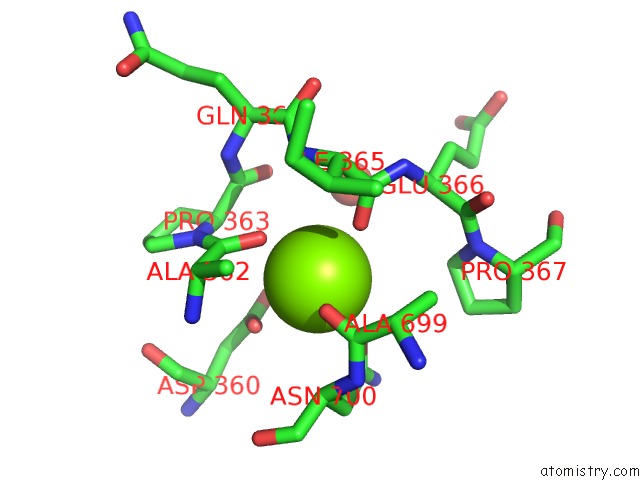
Mono view
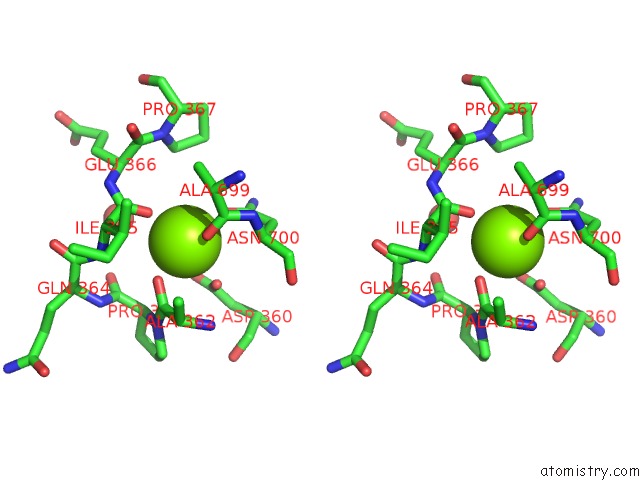
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5 within 5.0Å range:
|
Magnesium binding site 2 out of 4 in 6ubz
Go back to
Magnesium binding site 2 out
of 4 in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5
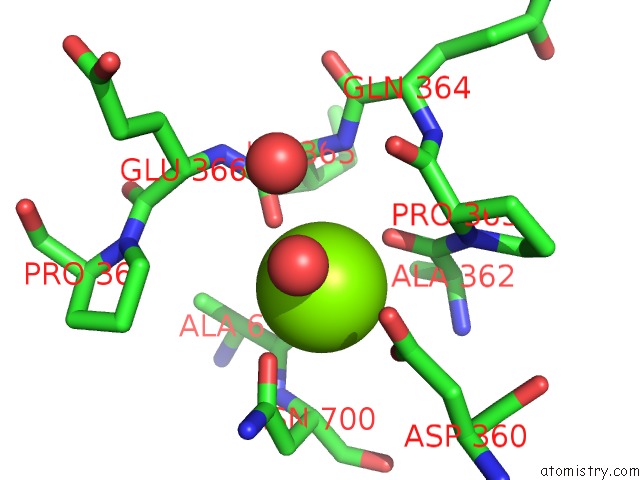
Mono view
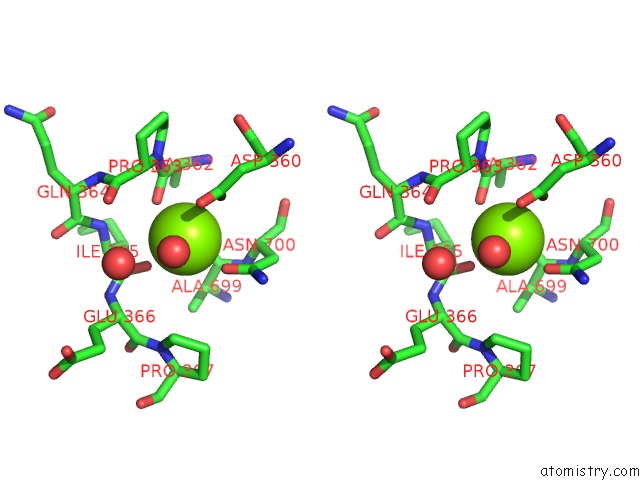
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5 within 5.0Å range:
|
Magnesium binding site 3 out of 4 in 6ubz
Go back to
Magnesium binding site 3 out
of 4 in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5

Mono view
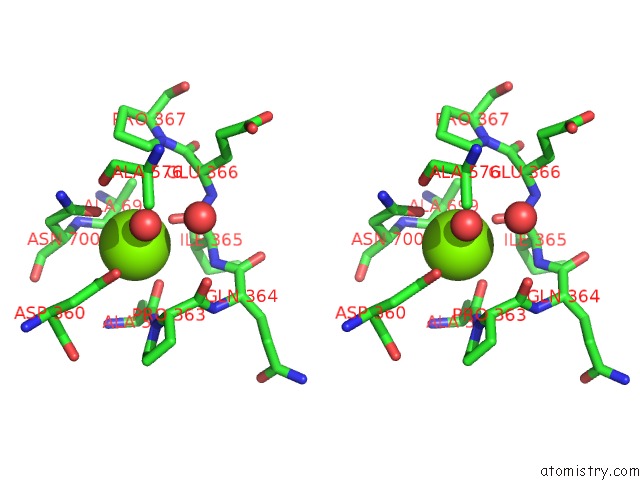
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 3 of Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5 within 5.0Å range:
|
Magnesium binding site 4 out of 4 in 6ubz
Go back to
Magnesium binding site 4 out
of 4 in the Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5
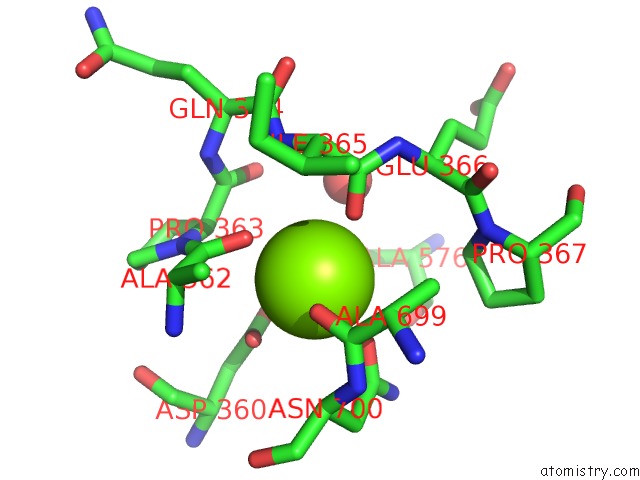
Mono view
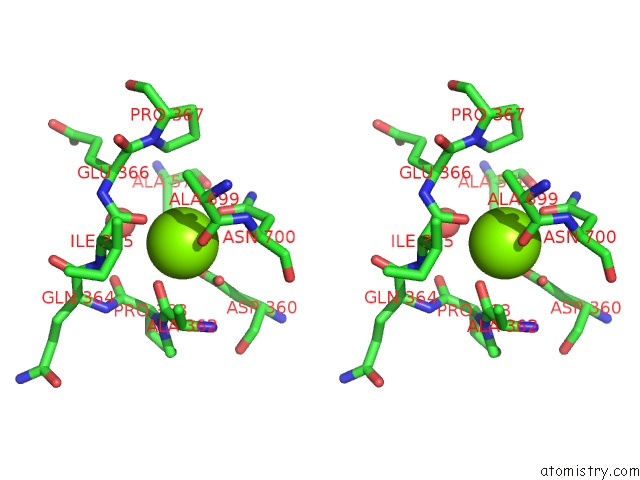
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 4 of Crystal Structure of D678A Goxa Bound to Glycine at pH 5.5 within 5.0Å range:
|
Reference:
K.J.Mamounis,
D.Avalos,
E.T.Yukl,
V.L.Davidson.
Kinetic and Structural Evidence That Asp-678 Plays Multiple Roles in Catalysis By the Quinoprotein Glycine Oxidase. J.Biol.Chem. V. 294 17463 2019.
ISSN: ESSN 1083-351X
PubMed: 31615898
DOI: 10.1074/JBC.RA119.011255
Page generated: Tue Oct 1 20:54:07 2024
ISSN: ESSN 1083-351X
PubMed: 31615898
DOI: 10.1074/JBC.RA119.011255
Last articles
Cl in 5X4NCl in 5X4P
Cl in 5X2P
Cl in 5X4I
Cl in 5X4J
Cl in 5X2S
Cl in 5X2T
Cl in 5X2O
Cl in 5X2N
Cl in 5X2M