Magnesium »
PDB 6wid-6wpj »
6wp5 »
Magnesium in PDB 6wp5: Pyruvate Kinase M2 Mutant-S37D
Enzymatic activity of Pyruvate Kinase M2 Mutant-S37D
All present enzymatic activity of Pyruvate Kinase M2 Mutant-S37D:
2.7.1.40;
2.7.1.40;
Protein crystallography data
The structure of Pyruvate Kinase M2 Mutant-S37D, PDB code: 6wp5
was solved by
S.Nandi,
M.Razzaghi,
D.Srivastava,
M.Dey,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.84 / 2.17 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 94.625, 117.567, 109.965, 90.00, 112.63, 90.00 |
| R / Rfree (%) | 21.7 / 26 |
Other elements in 6wp5:
The structure of Pyruvate Kinase M2 Mutant-S37D also contains other interesting chemical elements:
| Potassium | (K) | 4 atoms |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Pyruvate Kinase M2 Mutant-S37D
(pdb code 6wp5). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 3 binding sites of Magnesium where determined in the Pyruvate Kinase M2 Mutant-S37D, PDB code: 6wp5:
Jump to Magnesium binding site number: 1; 2; 3;
In total 3 binding sites of Magnesium where determined in the Pyruvate Kinase M2 Mutant-S37D, PDB code: 6wp5:
Jump to Magnesium binding site number: 1; 2; 3;
Magnesium binding site 1 out of 3 in 6wp5
Go back to
Magnesium binding site 1 out
of 3 in the Pyruvate Kinase M2 Mutant-S37D
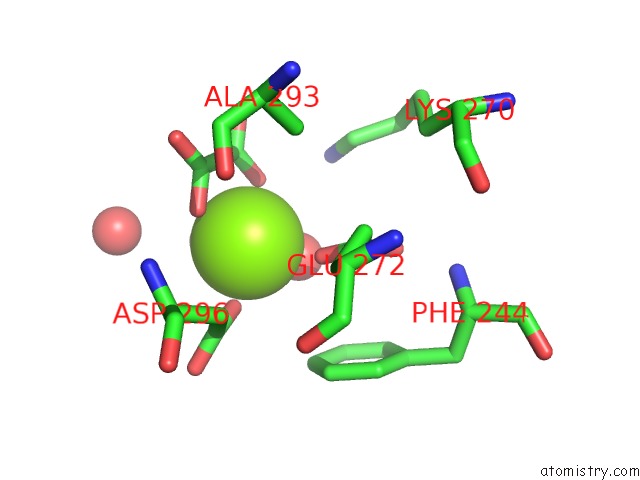
Mono view
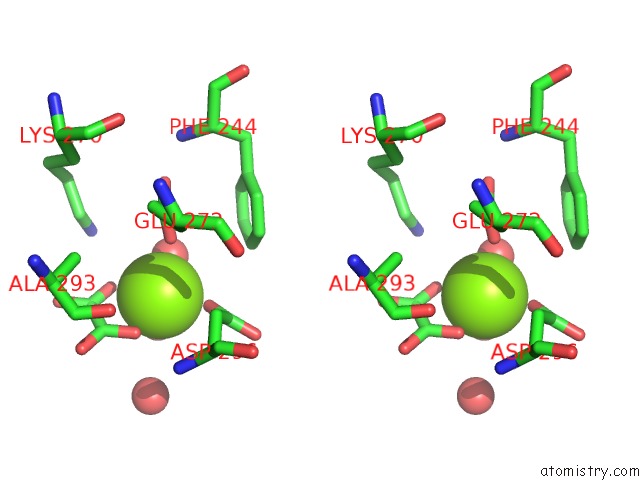
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Pyruvate Kinase M2 Mutant-S37D within 5.0Å range:
|
Magnesium binding site 2 out of 3 in 6wp5
Go back to
Magnesium binding site 2 out
of 3 in the Pyruvate Kinase M2 Mutant-S37D
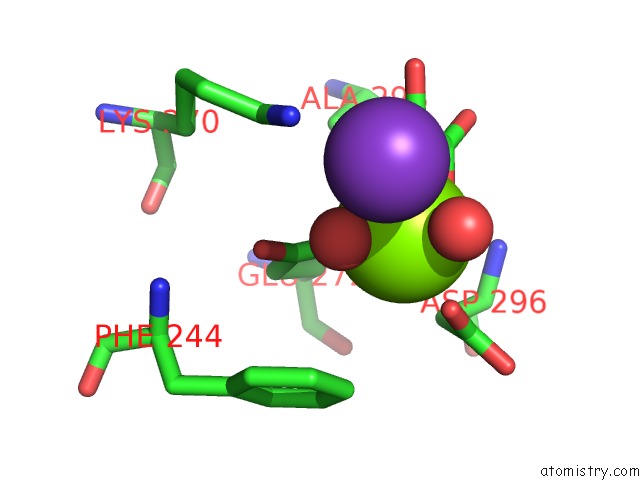
Mono view
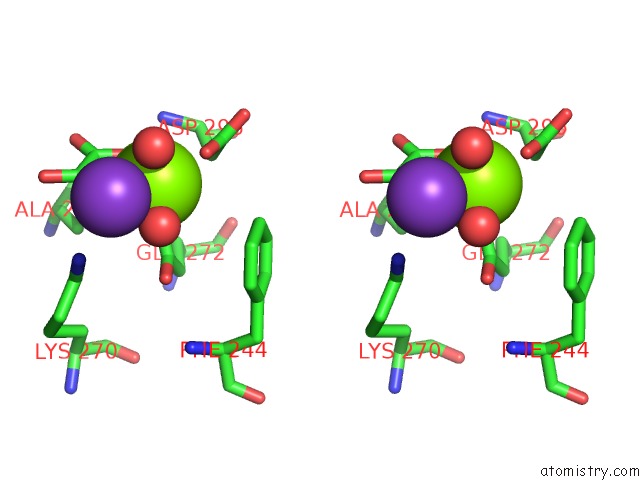
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Pyruvate Kinase M2 Mutant-S37D within 5.0Å range:
|
Magnesium binding site 3 out of 3 in 6wp5
Go back to
Magnesium binding site 3 out
of 3 in the Pyruvate Kinase M2 Mutant-S37D
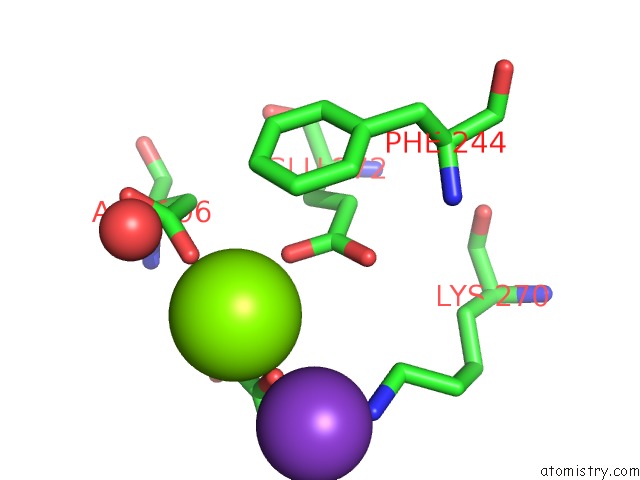
Mono view
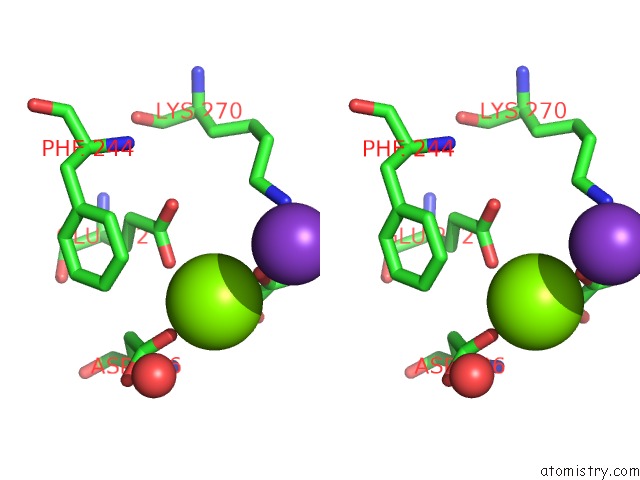
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 3 of Pyruvate Kinase M2 Mutant-S37D within 5.0Å range:
|
Reference:
S.Nandi,
M.Razzaghi,
D.Srivastava,
M.Dey.
Structural Basis For Allosteric Regulation of Pyruvate Kinase M2 By Phosphorylation and Acetylation. J.Biol.Chem. 2020.
ISSN: ESSN 1083-351X
PubMed: 32989054
DOI: 10.1074/JBC.RA120.015800
Page generated: Tue Oct 1 23:06:45 2024
ISSN: ESSN 1083-351X
PubMed: 32989054
DOI: 10.1074/JBC.RA120.015800
Last articles
Fe in 2YXOFe in 2YRS
Fe in 2YXC
Fe in 2YNM
Fe in 2YVJ
Fe in 2YP1
Fe in 2YU2
Fe in 2YU1
Fe in 2YQB
Fe in 2YOO