Magnesium »
PDB 6zup-7a1x »
7a0c »
Magnesium in PDB 7a0c: X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
Protein crystallography data
The structure of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn, PDB code: 7a0c
was solved by
C.Cavazza,
S.Menage,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 43.88 / 1.90 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 86.809, 93.796, 124.222, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.2 / 22.2 |
Other elements in 7a0c:
The structure of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn also contains other interesting chemical elements:
| Iron | (Fe) | 2 atoms |
| Chlorine | (Cl) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
(pdb code 7a0c). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 5 binding sites of Magnesium where determined in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn, PDB code: 7a0c:
Jump to Magnesium binding site number: 1; 2; 3; 4; 5;
In total 5 binding sites of Magnesium where determined in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn, PDB code: 7a0c:
Jump to Magnesium binding site number: 1; 2; 3; 4; 5;
Magnesium binding site 1 out of 5 in 7a0c
Go back to
Magnesium binding site 1 out
of 5 in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
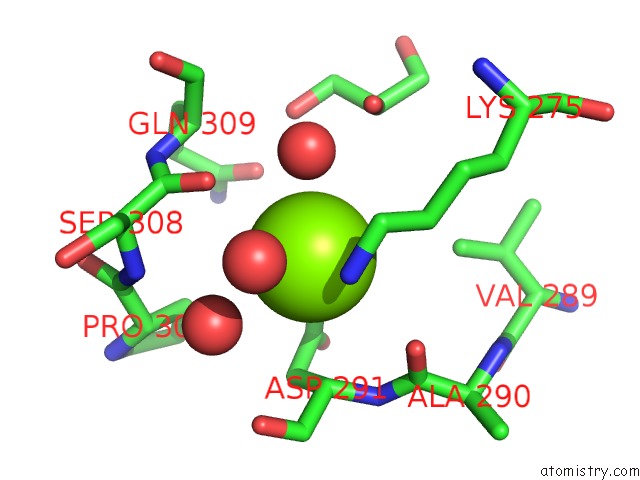
Mono view
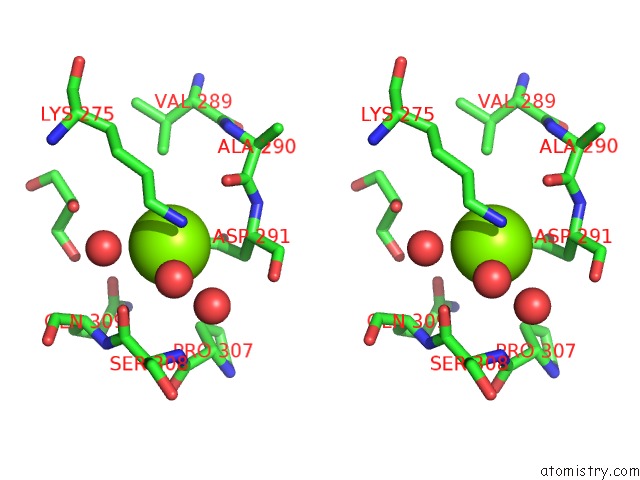
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn within 5.0Å range:
|
Magnesium binding site 2 out of 5 in 7a0c
Go back to
Magnesium binding site 2 out
of 5 in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
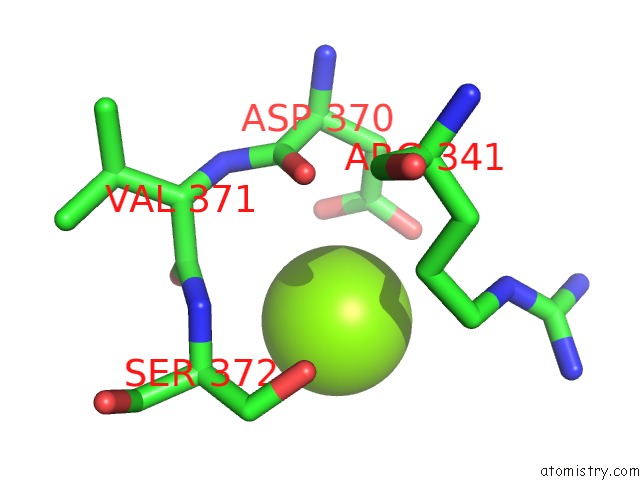
Mono view
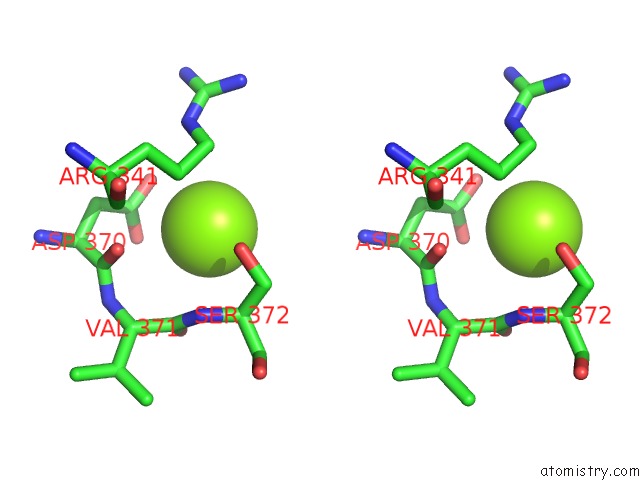
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn within 5.0Å range:
|
Magnesium binding site 3 out of 5 in 7a0c
Go back to
Magnesium binding site 3 out
of 5 in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
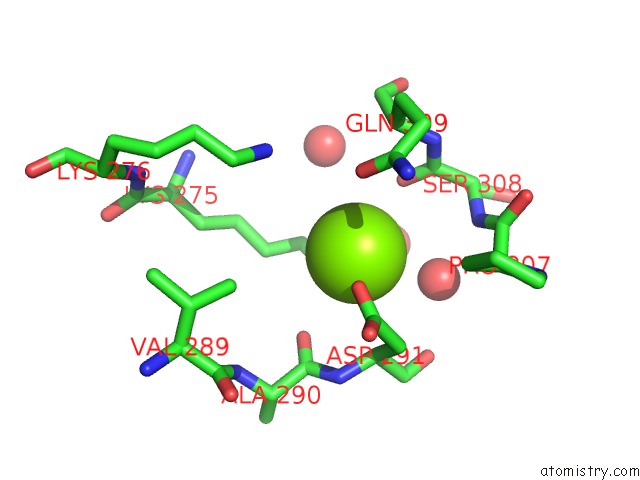
Mono view
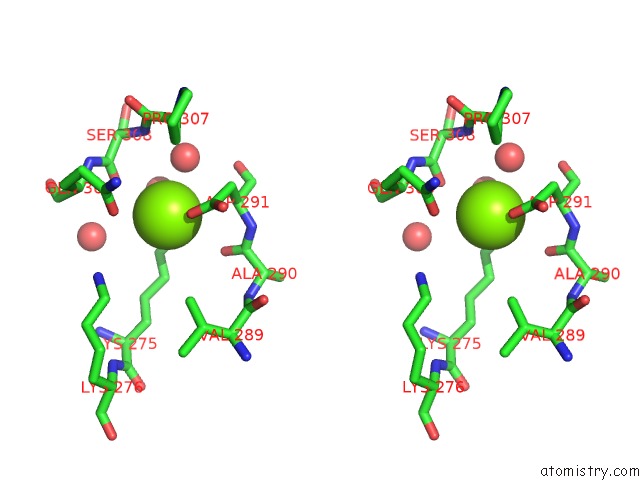
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 3 of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn within 5.0Å range:
|
Magnesium binding site 4 out of 5 in 7a0c
Go back to
Magnesium binding site 4 out
of 5 in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
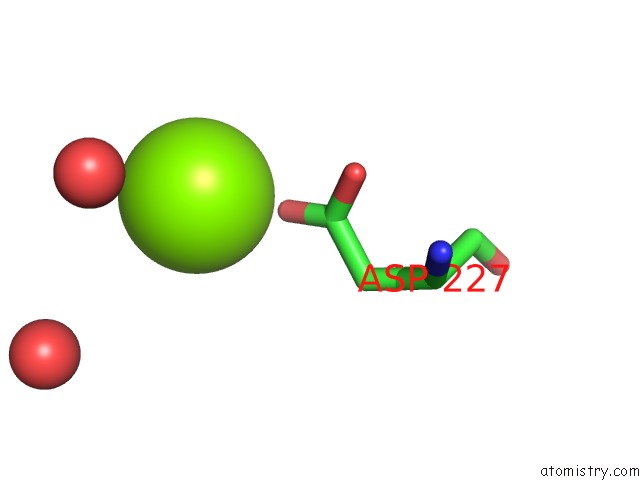
Mono view
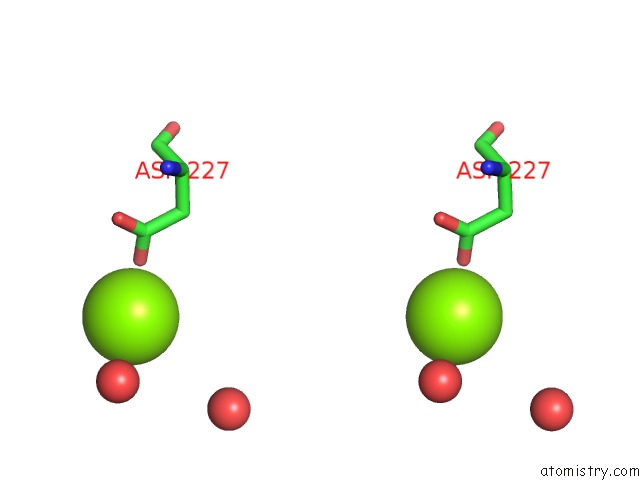
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 4 of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn within 5.0Å range:
|
Magnesium binding site 5 out of 5 in 7a0c
Go back to
Magnesium binding site 5 out
of 5 in the X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn
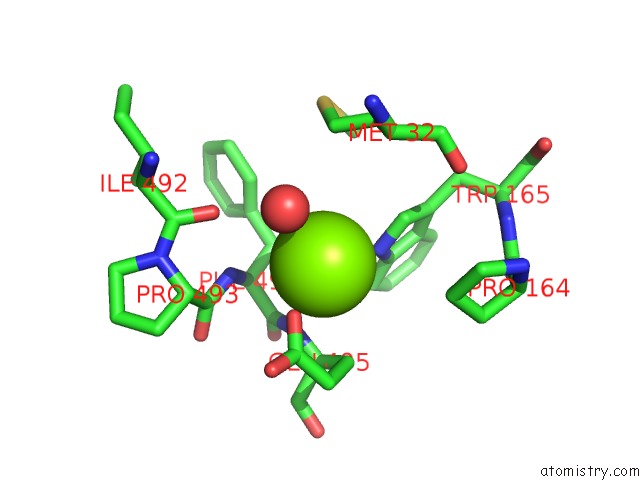
Mono view
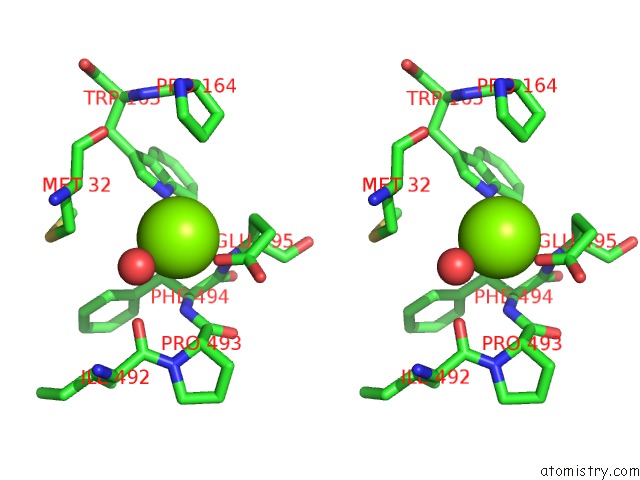
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 5 of X-Ray Structure of Nika From Escherichia Coli in Complex with Fe-6- ME2-Bpmcn within 5.0Å range:
|
Reference:
S.Lopez,
C.Marchi-Delapierre,
C.Cavazza,
S.Menage.
A Selective Sulfide Oxidation Catalyzed By Heterogeneous Artificial Metalloenzymes Iron@Nika. Chemistry 2020.
ISSN: ISSN 0947-6539
PubMed: 33079395
DOI: 10.1002/CHEM.202003746
Page generated: Wed Oct 2 03:22:05 2024
ISSN: ISSN 0947-6539
PubMed: 33079395
DOI: 10.1002/CHEM.202003746
Last articles
F in 7JHWF in 7JHD
F in 7I18
F in 7I2F
F in 7I2M
F in 7I2A
F in 7I2D
F in 7HNS
F in 7HOG
F in 7HO4