Magnesium »
PDB 8aom-8auy »
8asg »
Magnesium in PDB 8asg: Structure of the Sftsv L Protein Bound in A Resting State [Resting]
Enzymatic activity of Structure of the Sftsv L Protein Bound in A Resting State [Resting]
All present enzymatic activity of Structure of the Sftsv L Protein Bound in A Resting State [Resting]:
2.7.7.48;
2.7.7.48;
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Structure of the Sftsv L Protein Bound in A Resting State [Resting]
(pdb code 8asg). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Structure of the Sftsv L Protein Bound in A Resting State [Resting], PDB code: 8asg:
In total only one binding site of Magnesium was determined in the Structure of the Sftsv L Protein Bound in A Resting State [Resting], PDB code: 8asg:
Magnesium binding site 1 out of 1 in 8asg
Go back to
Magnesium binding site 1 out
of 1 in the Structure of the Sftsv L Protein Bound in A Resting State [Resting]

Mono view
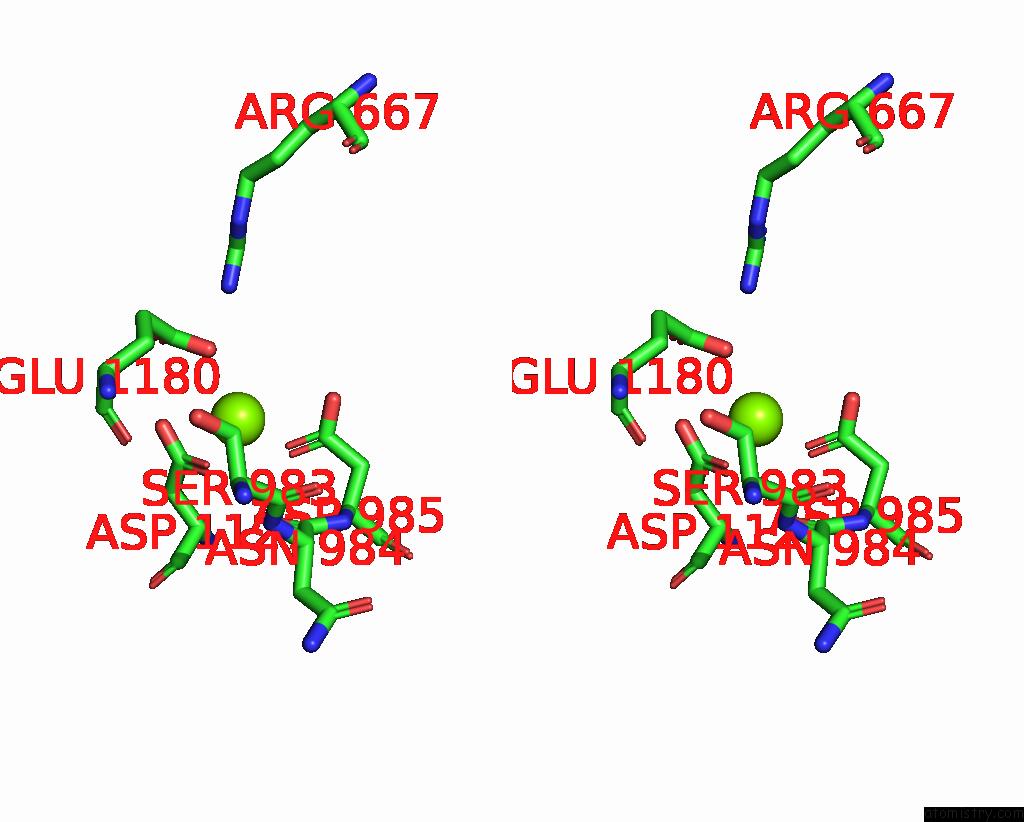
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Structure of the Sftsv L Protein Bound in A Resting State [Resting] within 5.0Å range:
|
Reference:
H.M.Williams,
S.R.Thorkelsson,
D.Vogel,
M.Milewski,
C.Busch,
S.Cusack,
K.Grunewald,
E.R.J.Quemin,
M.Rosenthal.
Structural Insights Into Viral Genome Replication By the Severe Fever with Thrombocytopenia Syndrome Virus L Protein Nucleic Acids Res. 2023.
ISSN: ESSN 1362-4962
DOI: 10.1093/NAR/GKAC1249
Page generated: Thu Oct 3 18:43:53 2024
ISSN: ESSN 1362-4962
DOI: 10.1093/NAR/GKAC1249
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1