Magnesium »
PDB 9mh0-9nea »
9mhg »
Magnesium in PDB 9mhg: Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation
Enzymatic activity of Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation
All present enzymatic activity of Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation:
2.7.1.137; 2.7.11.1; 3.6.5.2;
2.7.1.137; 2.7.11.1; 3.6.5.2;
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation
(pdb code 9mhg). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation, PDB code: 9mhg:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation, PDB code: 9mhg:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 9mhg
Go back to
Magnesium binding site 1 out
of 2 in the Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation
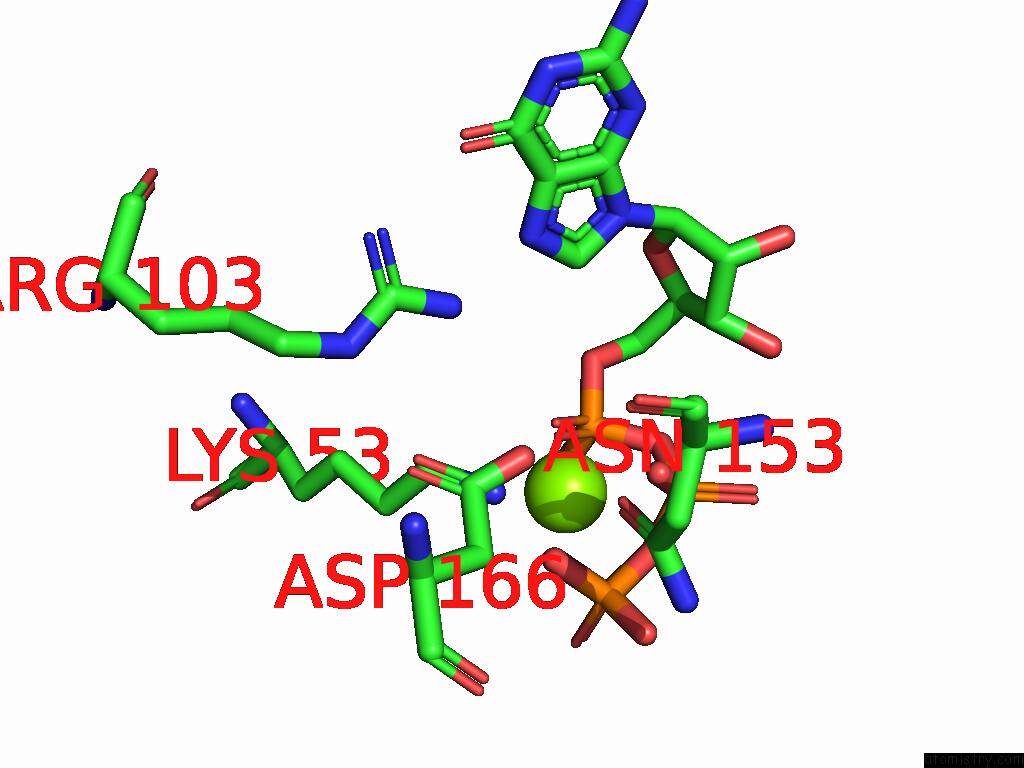
Mono view
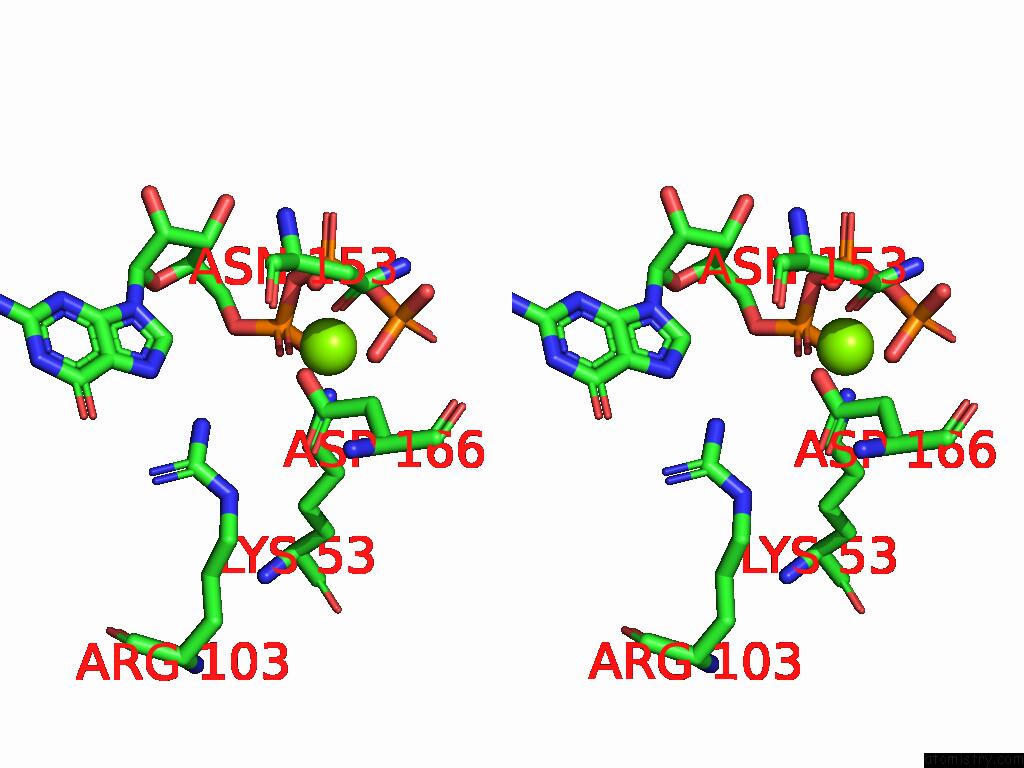
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 9mhg
Go back to
Magnesium binding site 2 out
of 2 in the Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation
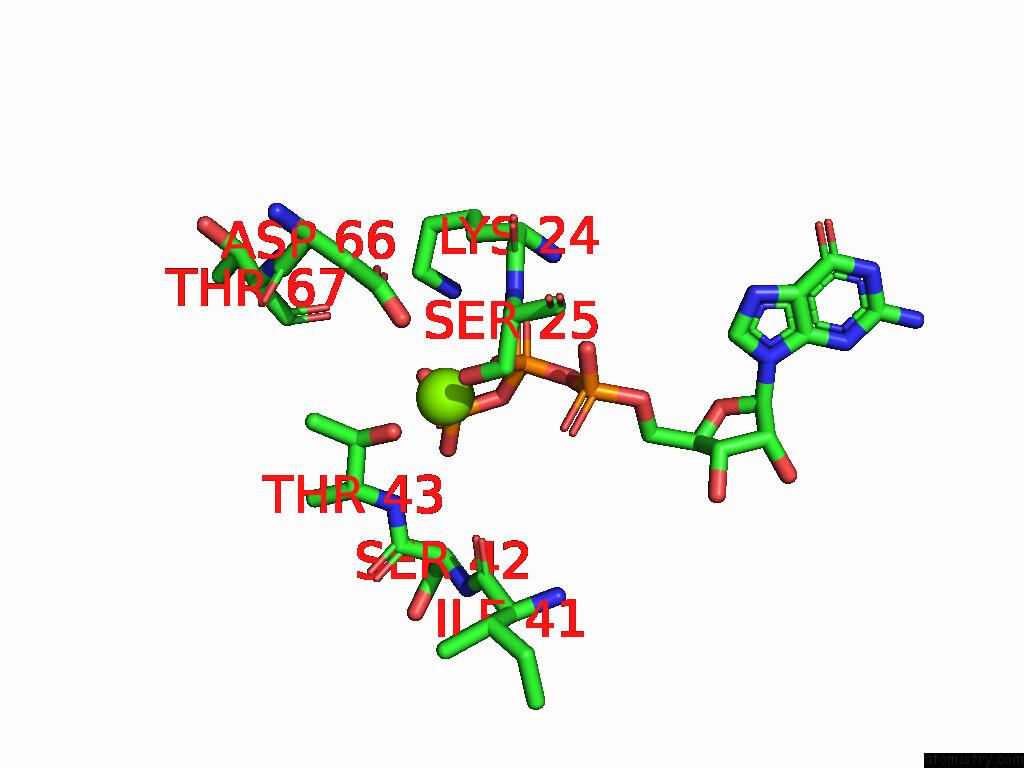
Mono view
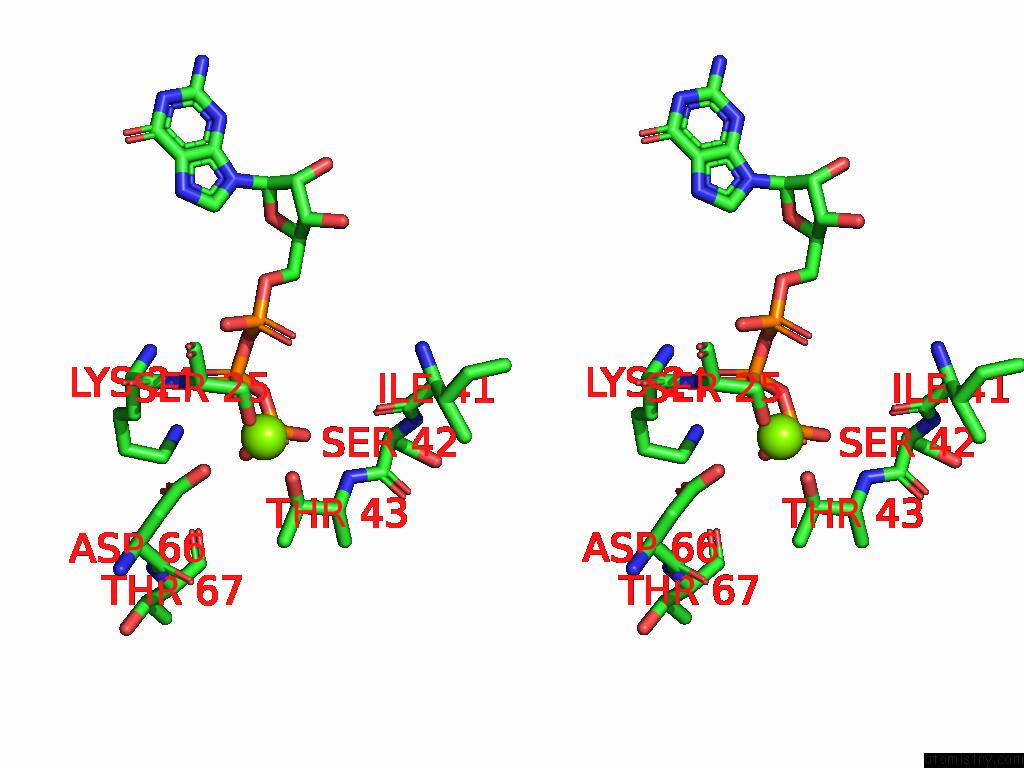
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Cryo Em Reconstruction of PI3KC3-C1 in Complex with Human RAB1A(Q70L), VPS34 Kinase Domain in the Inactive Conformation within 5.0Å range:
|
Reference:
A.S.I.Cook,
M.Chen,
T.N.Ngyuen,
A.Claveras-Cabezudo,
G.Khuu,
S.Rao,
S.N.Garcia,
M.Yang,
A.T.Iavarone,
X.Ren,
M.Lazarou,
G.Hummer,
J.H.Hurley.
Structural Pathway For Class III Pi 3-Kinase Activation By the Myristoylated Gtpbinding Pseudokinase VPS15 To Be Published.
Page generated: Sat Aug 16 06:25:06 2025
Last articles
Mo in 7WY2Mo in 7WY1
Mo in 7W9J
Mo in 7W9D
Mo in 7UTA
Mo in 7UT9
Mo in 7UT8
Mo in 7UT7
Mo in 7UT6
Mo in 7PX0