Magnesium »
PDB 5l5v-5lcf »
5l9n »
Magnesium in PDB 5l9n: Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp
Protein crystallography data
The structure of Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp, PDB code: 5l9n
was solved by
V.Rubio,
C.Palanca,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 28.10 / 1.90 |
| Space group | I 21 3 |
| Cell size a, b, c (Å), α, β, γ (°) | 88.868, 88.868, 88.868, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.4 / 22.3 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp
(pdb code 5l9n). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp, PDB code: 5l9n:
In total only one binding site of Magnesium was determined in the Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp, PDB code: 5l9n:
Magnesium binding site 1 out of 1 in 5l9n
Go back to
Magnesium binding site 1 out
of 1 in the Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp
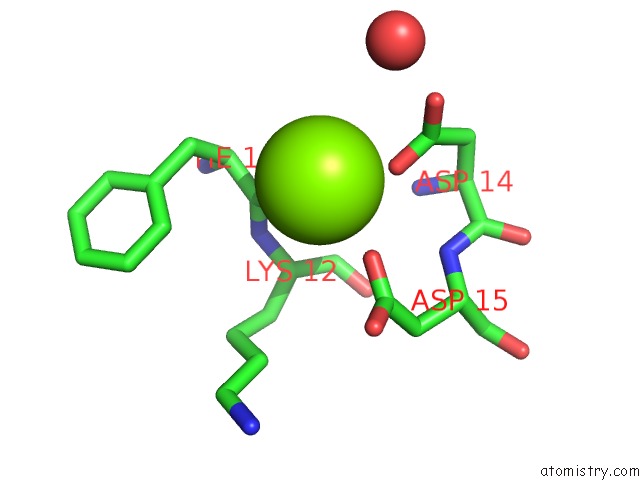
Mono view
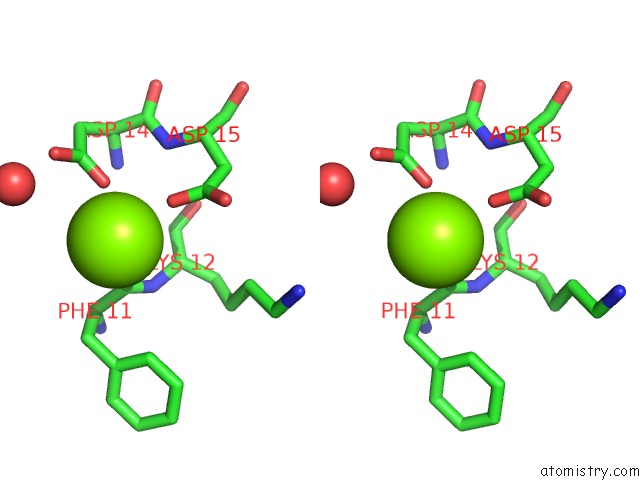
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Structure of Uridylylated Glnb From Escherichia Coli Bound to Atp within 5.0Å range:
|
Reference:
C.Palanca,
V.Rubio.
Effects of T-Loop Modification on the Pii-Signalling Protein: Structure of Uridylylated Escherichia Coli Glnb Bound to Atp. Environ Microbiol Rep V. 9 290 2017.
ISSN: ESSN 1758-2229
PubMed: 28345298
DOI: 10.1111/1758-2229.12533
Page generated: Tue Aug 12 13:44:58 2025
ISSN: ESSN 1758-2229
PubMed: 28345298
DOI: 10.1111/1758-2229.12533
Last articles
Mg in 6JNXMg in 6JMG
Mg in 6JNL
Mg in 6JLV
Mg in 6JLN
Mg in 6JLL
Mg in 6JLK
Mg in 6JLM
Mg in 6JLJ
Mg in 6JIL