Magnesium »
PDB 5tff-5tv8 »
5tt5 »
Magnesium in PDB 5tt5: Escherichia Coli Liga (K115M) in Complex with Nad+
Enzymatic activity of Escherichia Coli Liga (K115M) in Complex with Nad+
All present enzymatic activity of Escherichia Coli Liga (K115M) in Complex with Nad+:
6.5.1.2;
6.5.1.2;
Protein crystallography data
The structure of Escherichia Coli Liga (K115M) in Complex with Nad+, PDB code: 5tt5
was solved by
Y.Goldgur,
M.-C.Unciuleac,
S.H.Shuman,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.48 / 1.55 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 40.907, 94.312, 150.927, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.7 / 20.5 |
Other elements in 5tt5:
The structure of Escherichia Coli Liga (K115M) in Complex with Nad+ also contains other interesting chemical elements:
| Zinc | (Zn) | 1 atom |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Escherichia Coli Liga (K115M) in Complex with Nad+
(pdb code 5tt5). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Escherichia Coli Liga (K115M) in Complex with Nad+, PDB code: 5tt5:
In total only one binding site of Magnesium was determined in the Escherichia Coli Liga (K115M) in Complex with Nad+, PDB code: 5tt5:
Magnesium binding site 1 out of 1 in 5tt5
Go back to
Magnesium binding site 1 out
of 1 in the Escherichia Coli Liga (K115M) in Complex with Nad+
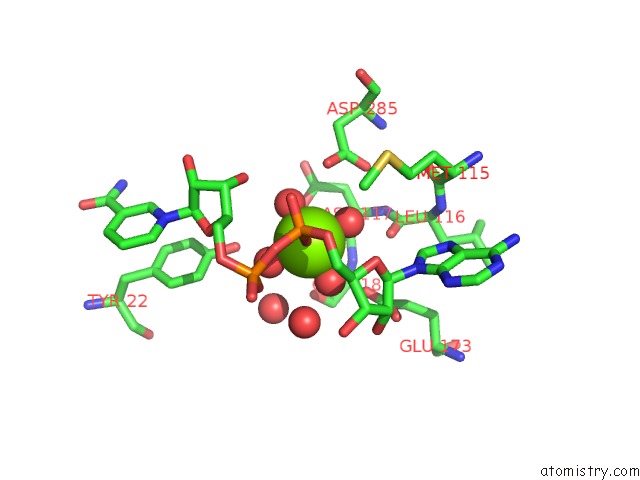
Mono view
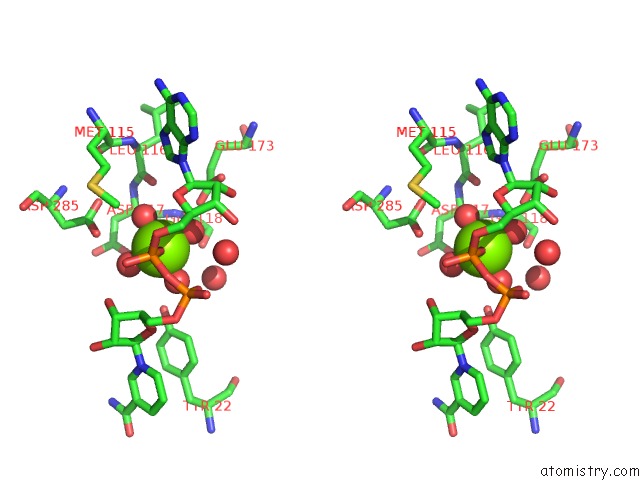
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Escherichia Coli Liga (K115M) in Complex with Nad+ within 5.0Å range:
|
Reference:
M.C.Unciuleac,
Y.Goldgur,
S.Shuman.
Two-Metal Versus One-Metal Mechanisms of Lysine Adenylylation By Atp-Dependent and Nad(+)-Dependent Polynucleotide Ligases. Proc. Natl. Acad. Sci. V. 114 2592 2017U.S.A..
ISSN: ESSN 1091-6490
PubMed: 28223499
DOI: 10.1073/PNAS.1619220114
Page generated: Tue Aug 12 20:33:26 2025
ISSN: ESSN 1091-6490
PubMed: 28223499
DOI: 10.1073/PNAS.1619220114
Last articles
Mn in 3DT2Mn in 3DJ8
Mn in 3DKY
Mn in 3DKX
Mn in 3DIY
Mn in 3D4K
Mn in 3DC6
Mn in 3DC5
Mn in 3DBN
Mn in 3CEV