Magnesium »
PDB 7r8p-7rf0 »
7rd8 »
Magnesium in PDB 7rd8: Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State
Enzymatic activity of Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State
All present enzymatic activity of Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State:
7.6.2.1;
7.6.2.1;
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State
(pdb code 7rd8). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State, PDB code: 7rd8:
In total only one binding site of Magnesium was determined in the Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State, PDB code: 7rd8:
Magnesium binding site 1 out of 1 in 7rd8
Go back to
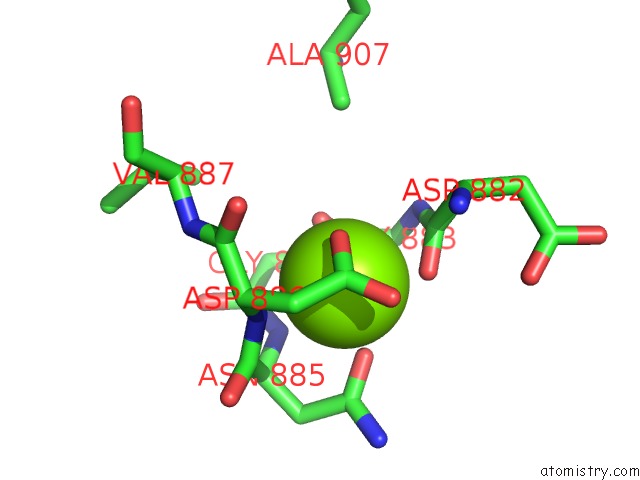
Magnesium binding site 1 out
of 1 in the Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E1-Atp State within 5.0Å range:
|
Reference:
L.Bai,
B.K.Jain,
Q.You,
H.D.Duan,
T.R.Graham,
H.Li.
Structure of the S. Cerevisiae P4B Atpase Lipid Flippase in the E2P State To Be Published.
Page generated: Thu Aug 14 14:59:20 2025
Last articles
Cl in 7HSYCl in 7HU9
Cl in 7HSQ
Cl in 7HRG
Cl in 7HRE
Cl in 7HRF
Cl in 7HR4
Cl in 7HQT
Cd in 9J00
Cd in 9J01