Magnesium »
PDB 6at2-6b5e »
6au9 »
Magnesium in PDB 6au9: Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid
Enzymatic activity of Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid
All present enzymatic activity of Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid:
2.3.3.9;
2.3.3.9;
Protein crystallography data
The structure of Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid, PDB code: 6au9
was solved by
I.V.Krieger,
J.C.Sacchettini,
Tb Structural Genomics Consortium (Tbsgc),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 49.49 / 2.10 |
| Space group | P 43 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 78.071, 78.071, 223.463, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16.3 / 19.9 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid
(pdb code 6au9). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid, PDB code: 6au9:
In total only one binding site of Magnesium was determined in the Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid, PDB code: 6au9:
Magnesium binding site 1 out of 1 in 6au9
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid
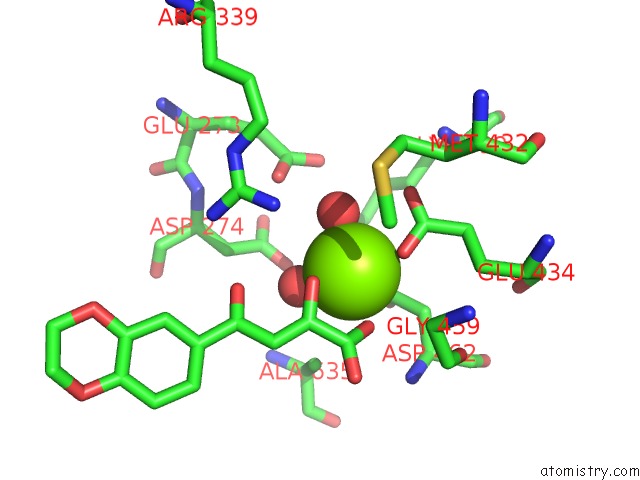
Mono view
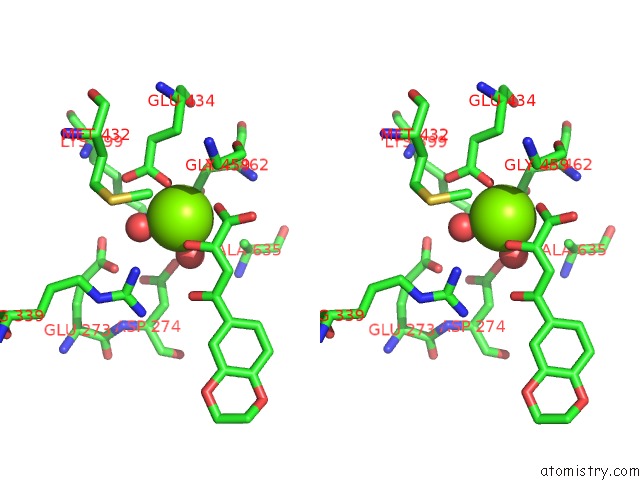
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of Mycobacterium Tuberculosis Malate Synthase in Complex with Dioxine-Phenyldiketoacid within 5.0Å range:
|
Reference:
J.F.Ellenbarger,
I.V.Krieger,
H.L.Huang,
S.Gomez-Coca,
T.R.Ioerger,
J.C.Sacchettini,
S.E.Wheeler,
K.R.Dunbar.
Anion-Pi Interactions in Computer-Aided Drug Design: Modeling the Inhibition of Malate Synthase By Phenyl-Diketo Acids. J Chem Inf Model V. 58 2085 2018.
ISSN: ESSN 1549-960X
PubMed: 30137983
DOI: 10.1021/ACS.JCIM.8B00417
Page generated: Wed Aug 13 02:21:11 2025
ISSN: ESSN 1549-960X
PubMed: 30137983
DOI: 10.1021/ACS.JCIM.8B00417
Last articles
Mg in 6KOUMg in 6KPE
Mg in 6KOQ
Mg in 6KNZ
Mg in 6KOP
Mg in 6KON
Mg in 6KOO
Mg in 6KO6
Mg in 6KO1
Mg in 6KMA