Magnesium »
PDB 6wj2-6wqx »
6wmd »
Magnesium in PDB 6wmd: Human SUN2-KASH4 Complex
Protein crystallography data
The structure of Human SUN2-KASH4 Complex, PDB code: 6wmd
was solved by
V.E.Cruz,
T.U.Schwartz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 69.94 / 1.50 |
| Space group | P 63 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 80.757, 80.757, 173.909, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 16.7 / 18.4 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Human SUN2-KASH4 Complex
(pdb code 6wmd). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the Human SUN2-KASH4 Complex, PDB code: 6wmd:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the Human SUN2-KASH4 Complex, PDB code: 6wmd:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 6wmd
Go back to
Magnesium binding site 1 out
of 2 in the Human SUN2-KASH4 Complex
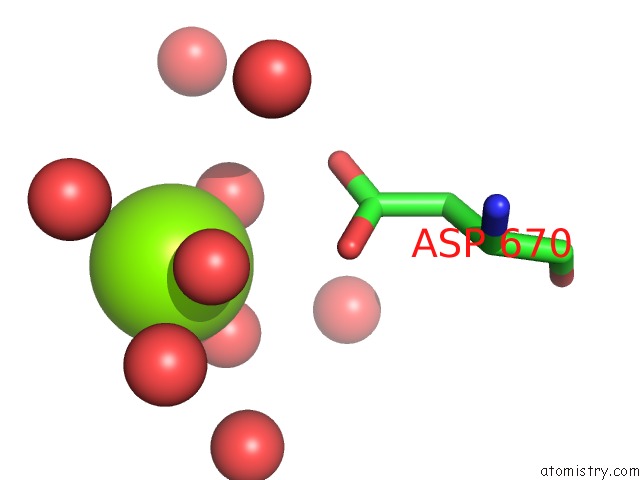
Mono view
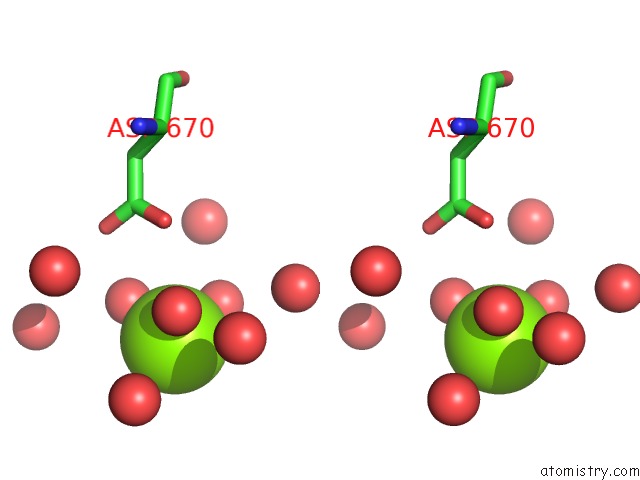
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Human SUN2-KASH4 Complex within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 6wmd
Go back to
Magnesium binding site 2 out
of 2 in the Human SUN2-KASH4 Complex
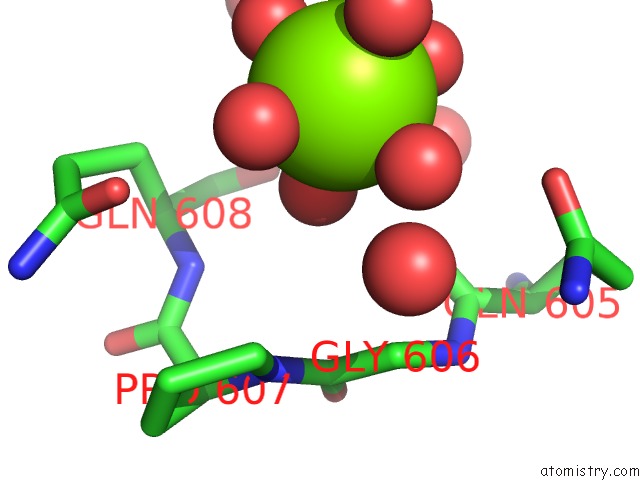
Mono view
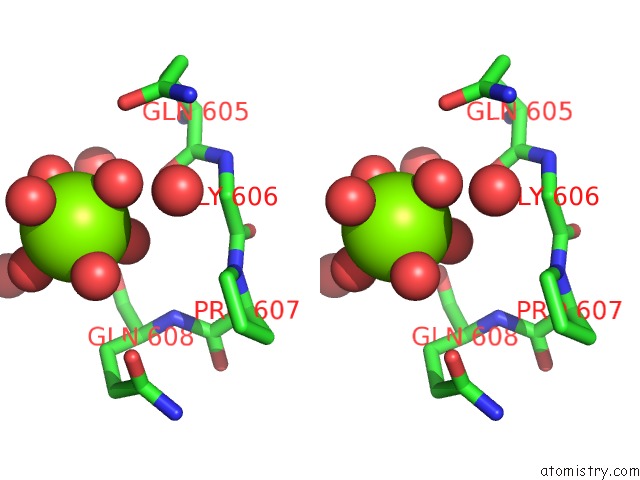
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Human SUN2-KASH4 Complex within 5.0Å range:
|
Reference:
V.E.Cruz,
F.Esra Demircioglu,
T.U.Schwartz.
Structural Analysis of Different Linc Complexes Reveals Distinct Binding Modes. J.Mol.Biol. V. 432 6028 2020.
ISSN: ESSN 1089-8638
PubMed: 33058875
DOI: 10.1016/J.JMB.2020.09.019
Page generated: Wed Aug 13 19:48:13 2025
ISSN: ESSN 1089-8638
PubMed: 33058875
DOI: 10.1016/J.JMB.2020.09.019
Last articles
Mg in 7E2WMg in 7E2B
Mg in 7E21
Mg in 7E0K
Mg in 7E20
Mg in 7E1Z
Mg in 7E1T
Mg in 7E1H
Mg in 7E1G
Mg in 7E12