Magnesium »
PDB 6j7r-6jlj »
6jdv »
Magnesium in PDB 6jdv: Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State
Protein crystallography data
The structure of Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State, PDB code: 6jdv
was solved by
W.Sun,
J.Yang,
Z.Cheng,
C.Liu,
K.Wang,
X.Huang,
Y.Wang,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 41.96 / 3.10 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 130.541, 158.340, 118.253, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 21.2 / 23.1 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State
(pdb code 6jdv). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total only one binding site of Magnesium was determined in the Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State, PDB code: 6jdv:
In total only one binding site of Magnesium was determined in the Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State, PDB code: 6jdv:
Magnesium binding site 1 out of 1 in 6jdv
Go back to
Magnesium binding site 1 out
of 1 in the Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State
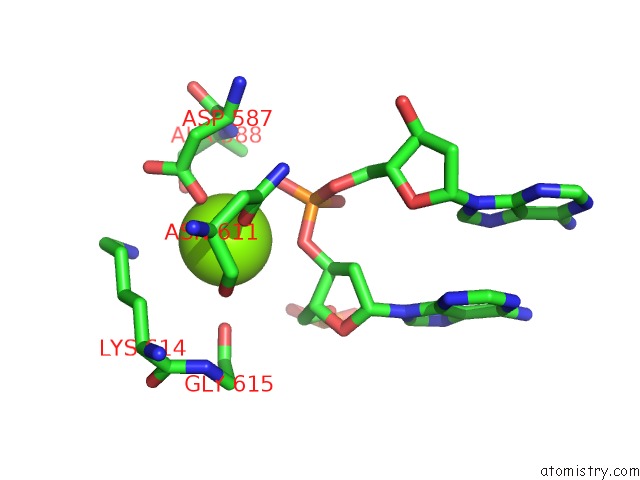
Mono view
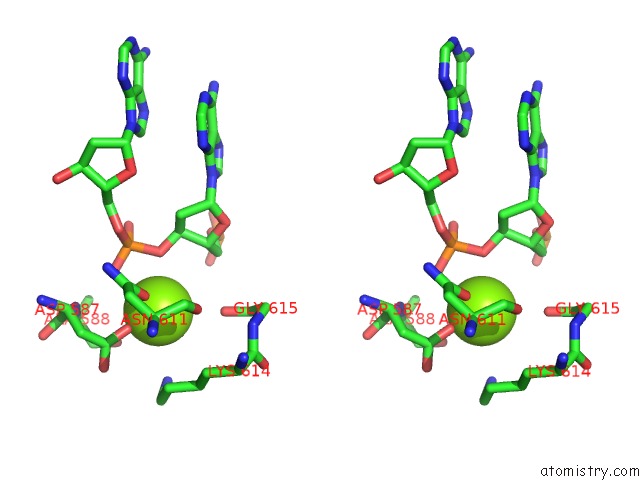
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Crystal Structure of NME1CAS9 in Complex with Sgrna and Target Dna (Atatgatt Pam) in Catalytic State within 5.0Å range:
|
Reference:
W.Sun,
J.Yang,
Z.Cheng,
N.Amrani,
C.Liu,
K.Wang,
R.Ibraheim,
A.Edraki,
X.Huang,
M.Wang,
J.Wang,
L.Liu,
G.Sheng,
Y.Yang,
J.Lou,
E.J.Sontheimer,
Y.Wang.
Structures of Neisseria Meningitidis CAS9 Complexes in Catalytically Poised and Anti-Crispr-Inhibited States. Mol.Cell 2019.
ISSN: ISSN 1097-2765
PubMed: 31668930
DOI: 10.1016/J.MOLCEL.2019.09.025
Page generated: Wed Aug 13 09:12:06 2025
ISSN: ISSN 1097-2765
PubMed: 31668930
DOI: 10.1016/J.MOLCEL.2019.09.025
Last articles
Mg in 7A17Mg in 7A0V
Mg in 6ZZX
Mg in 6ZXS
Mg in 7A0Q
Mg in 7A0P
Mg in 7A0C
Mg in 721P
Mg in 6ZXH
Mg in 6ZXG