Magnesium »
PDB 6ggi-6gog »
6gmo »
Magnesium in PDB 6gmo: Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm)
Enzymatic activity of Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm)
All present enzymatic activity of Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm):
6.3.2.2;
6.3.2.2;
Protein crystallography data
The structure of Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm), PDB code: 6gmo
was solved by
E.D.Lenherr,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.72 / 1.75 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 58.770, 109.860, 84.750, 90.00, 98.72, 90.00 |
| R / Rfree (%) | 16.3 / 19.8 |
Magnesium Binding Sites:
The binding sites of Magnesium atom in the Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm)
(pdb code 6gmo). This binding sites where shown within
5.0 Angstroms radius around Magnesium atom.
In total 2 binding sites of Magnesium where determined in the Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm), PDB code: 6gmo:
Jump to Magnesium binding site number: 1; 2;
In total 2 binding sites of Magnesium where determined in the Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm), PDB code: 6gmo:
Jump to Magnesium binding site number: 1; 2;
Magnesium binding site 1 out of 2 in 6gmo
Go back to
Magnesium binding site 1 out
of 2 in the Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm)
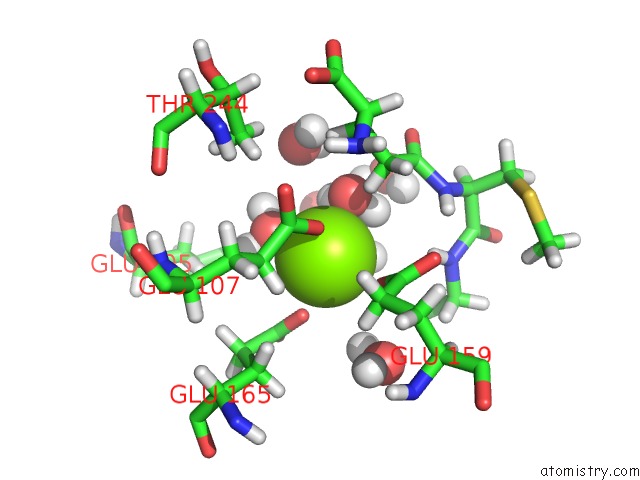
Mono view
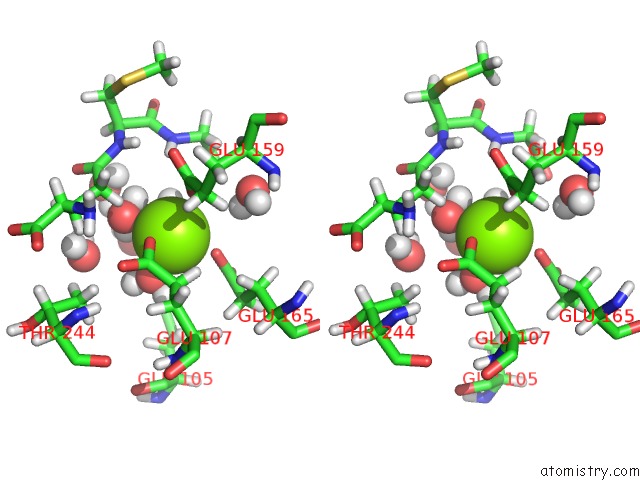
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 1 of Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm) within 5.0Å range:
|
Magnesium binding site 2 out of 2 in 6gmo
Go back to
Magnesium binding site 2 out
of 2 in the Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm)
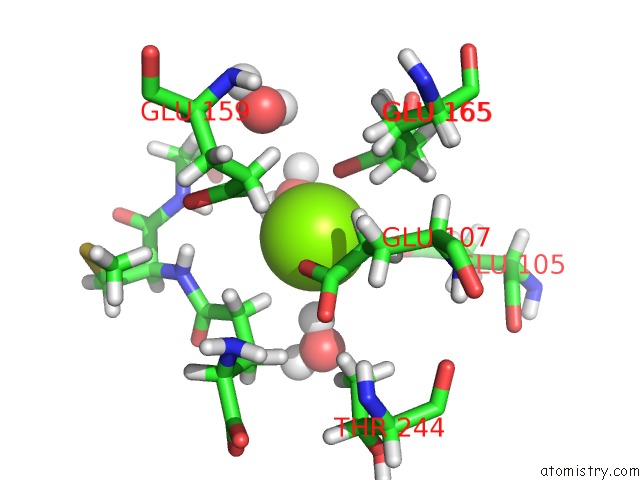
Mono view
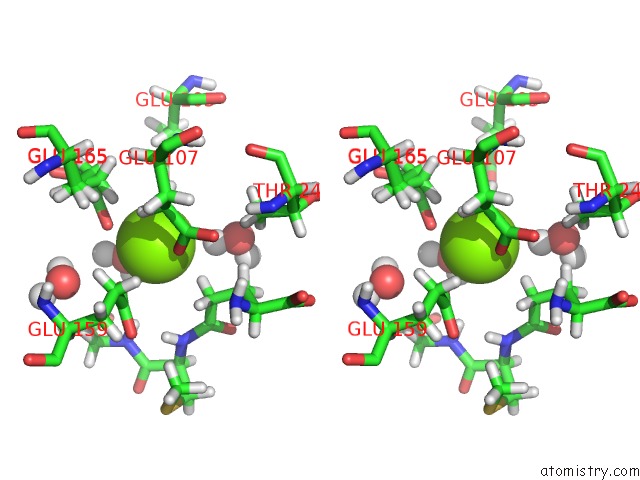
Stereo pair view

Mono view

Stereo pair view
A full contact list of Magnesium with other atoms in the Mg binding
site number 2 of Plant Glutamate Cysteine Ligase (Gcl) in Complex with Non-Reducing Gsh (Gsm) within 5.0Å range:
|
Reference:
Y.Yang,
E.Lenherr,
R.Gromes,
S.Wang,
M.Wirtz,
R.Hell,
K.Scheffzek,
T.Rausch.
Regulation of Plant Glutamylcysteine Ligase: Redox-Mediated Enzyme Activation Does Not Require Homodimerization To Be Published.
Page generated: Tue Oct 1 01:16:44 2024
Last articles
Mg in 5HPYMg in 5HPZ
Mg in 5HP1
Mg in 5HPO
Mg in 5HPH
Mg in 5HOO
Mg in 5HOK
Mg in 5HOB
Mg in 5HO5
Mg in 5HMQ